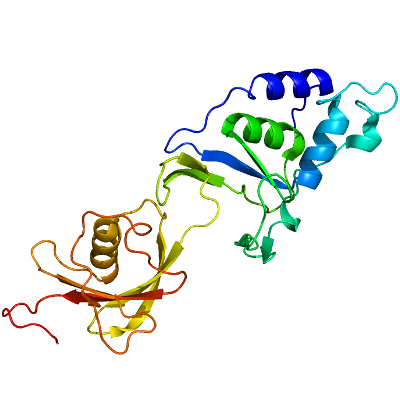
USP4
PDB:3JYU
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:BC125131
Entry Clone Source:MGC
SGC Clone Accession:usp04.001.229.048D10 (SDC048D10)
Tag:N-terminal tag: MGSSHHHHHHSSGLVPRGS
Host:
Construct
Prelude:
Sequence:mgsshhhhhhssglvprgsRERPDVETQKTELGALMGTTLQRGAQWYLIDSRWFKQWKKYVGFDSWDMYNVGEHNLFPGPIDNSGLFSDPESQTLKEHLIDELDYVLVPAEAWNKLLNWYGCVEGQQPIVRKVVEHGLFVKHCKVEVYLLELKLCENSDPTNVLSCHFSKADTIATIEKEMRKLFNIPAERETRLWNKYMSNTYE QLSKLDNTIQDAGLYQGQVLVIEPQNEDGTWPRQSL
Vector:pET28a-LIC
Growth
Medium:
Antibiotics:
Procedure:Competent BL21 (DE3) cells (Invitrogen C6000-03) were transformed and grown using the LEX system (Harbinger BEC) at 37 degC in 2L bottles (VWR 89000-242). In order to obtain the selenomethionyl derivative of the Usp4 domain, the cells were grown in M9 medium supplemented with supplemented with 150 mM glycerol, 100 µg/ml Kanamycin and 600 µl antifoam 204 (Sigma A-8311) using a M9 SeMET High-Yield growth media kit package (Medicilon MD045004-50L) according to manufacturer's instruction. When OD600 0.8 - 1.2 was reached, the temperature was reduced to 20 degC, and one hour later protein expression was induced with 100 µM IPTG (BioShop IPT001) and the culture was incubated overnight (16 hours) at 20 degC. Cell pellets were collected by centrifugation (12,227 x g, 20 min), frozen and stored at -80 degC.
Purification
Procedure
The cleared lysate was loaded onto a 3 mL TALON metal-affinity resin column (Clontech Laboratories 635504) at 4ºC. The column was washed with 10 mL Wash Buffer A, 10 mL Wash Buffer B and 10 mL Wash Buffer A. The protein was eluted with 6 mL Elution Buffer. The N-terminal His-tag was removed by overnight incubation of the protein with thrombin (1 unit/mg protein) at 4°C. The protein was finally purified by gel filtration on a HighLoad 16/60 Superdex 200 column (GE Healthcare 17-1069-01) using Gel Filtration Buffer. Fractions containing SeMet Usp4 were pooled and concentrated by ultrafiltration. The yield of the protein was 18 mg per liter of bacterial culture.
Extraction
Procedure
After resuspension in 30 mL per liter bacterial culture of Lysis Buffer, cells were lysed using a Microfluidics M110-EH microfluidizer at 18,000 psi.
Concentration:
Ligand
MassSpec:
Crystallization:Crystals of the Usp4.001.229 selenomethionyl derivative were grown at 291 K using the hanging drop method by mixing 1 volume of 9 mg/ml protein solution with 1 volume of a solution of 20% (w/v) PEG 4000, 10% (v/v) isopropanool, 0.1 M sodium HEPES, pH 7.2, and 0.001 M dithiothreitol in water and equilibrating the resulting drop against 1 M NaCl (well solution). The crystals were cryoprotected by immersion in the well solution mixed in 1:1 ratio with a water solution containing 60% (v/v) ethylene glycol and 200 mg/ml 3-(1-pyridino)-propane sulfonate (NDSB-201, Calbiochem 480005) and frozen by immersion in liquid nitrogen.
NMR Spectroscopy:
Data Collection:Diffraction data from a crystal of the selenomethionyl derivative of the Usp4 domain were collected at beamline APS 17-ID of the GM/CA-CAT at the Advanced Photon Source (Argonne National Laboratory). The data set was integrated and scaled using the MOSFLM and SCALA programs.
Data Processing:The structure was solved using a combination of MR and SAD techniques using the program PHASER and an NMR model (PDB entry 1W6V), to generate a mixed model (SCRWL server) for the final search model. The MR solution was then used to locate the Selenomethionine sites using phenix.autosol. Positions of selenomethionines and phases were additionally refined using SHARP. Initial model-building, density modification and refinement were performed using phenix.autobuild. Later stages of refinement and model building were performed through iterative model building using the graphics program Coot and phenix.refine, which led to a model with an R factor of 18.4 % (Rfree 23.3 %) for data between 41.2 and 2.37 Å.