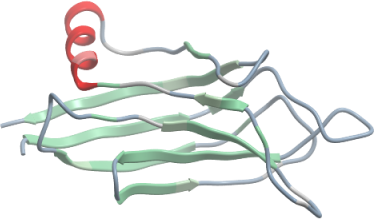
ITSN2
PDB:3JZY
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:BC146779
Entry Clone Source:OpenBiosystems EHS1001-99608273 (SGC 30-C2)
SGC Clone Accession:HPC098-C04
ITSN2:D1174-E1665
Tag:N-terminal tag: mhhhhhhssgrenlyfq*g
Host:BL21-CodonPlus(DE3)-V2R pRARE2
Construct
Prelude:Tag not removed.cDNA sequence verified by sequencing and it matches expected one.The cDNA template used contains a mutation K1525Q compared with reference sequence NP_062541.2.The construct contains 3 domains, only the C-terminal C2 domain was crystallized.
Sequence:mhhhhhhssgrenlyfqgDTMQPIERKRQGYIHELIQTEERYMADLQLVVEVFQKRMAESGFLTEGEMALIFVNWKELIMSNTKLLKALRVRKKTGGEKMPVQMIGDILAAELSHMQAYIRFCSCQLNGAALLQQKTDEDTDFKEFLKKLASDPRCKGMPLSSFLLKPMQRITRYPLLIRSILENTPESHADHSSLKLALERAEELCSQVNEGVREKENSDRLEWIQAHVQCEGLAEQLIFNSLTNCLGPRKLLHSGKLYKTKSNKELHGFLFNDFLLLTYMVKQFAVSSGSEKLFSSKSNAQFKMYKTPIFLNEVLVKLPTDPSSDEPVFHISHIDRVYTLRTDNINERTAWVQKIKAASEQYIDTEKKQREKAYQARSQKTSGIGRLMVHVIEATELKACKPNGKSNPYCEISMGSQSYTTRTIQDTLNPKWNFNCQFFIKDLYQDVLCLTLFDRDQFSPDDFLGRTEIPVAKIRTEQESKGPMTRRLLLHEVPTGEVWVRFDLQLFE
Vector:pET28-mhl (GI:134105571)
Growth
Medium:Terrific Broth
Antibiotics:Kanamycin 50 µg/mL + Chloramphenicol 25 µg/mL
Procedure:LEX Bubbling. The target protein was expressed in E. coli by inoculating 100 mL of overnight culture grown in Luria-Bertani medium into a 1.8 L of Terrific Broth medium in the presence of 50 µg/mL kanamycin and 25 µg/mL chloramphenicol at 37 degC. When OD600 reached ~3.0, the temperature of the medium was lowered to 15 degC and the culture was induced with 0.5 mM IPTG. The cells were allowed to grow overnight before they were harvested and flash frozen in liquid nitrogen and stored at -80 degC.
Purification
Procedure
The lysate was centrifuged at 16,000 rpm for 60 minutes and the supernatant was mixed with 4 mL 50% Ni-NTA beads, and incubated at 4 degC on roller drum for 1 hours. The supernatant was then passed through a gravity column (Poly-Prep, Bio-Rad, Catalog #731-1550) and the beads were washed using 50 mL binding buffer followed by 50mL washing buffer. The protein bound to beads were then eluted using 15 mL elution buffer. The flow-through was collected and loaded onto Supderdex-75 26/60 gel filtration column. Eluted fractions were pooled and concentrated using amicon centrifugal filter (m.w. cut-off 10,000 ). The purity of the proteins was higher than 95% judged by SDS-PAGE.
Extraction
Procedure
Frozen cells from 6L culture were thawed and resuspended in 400 mL extraction buffer with freshly added final concentration of 1mM PMSF/Benzamidine, 0.5% CHAPS and 5U/mL Benzonase (Sigma Catalog # E1014, 250U/µL), and supplemented with 1mL protease inhibitor cocktail (SIGMA Catalog # P8849), and lysed using sonication at 10 seconds 50% duty cycle for 5 minutes at 120W.
Concentration:23.43 mg/mL
Ligand
MassSpec:Native: 59018.16, expected 59013.88
Crystallization:Stock protein solution was added with a final concentration of 2 mM CaCl2 and crystallization was setup in sitting drops using Red Wings and SGC-I screens kits. No crystals were observed within one month of setup. Crystal used for data collection was harvested from a drop containing 20% PEG 3350, 0.2M CaCl2 in the well solution more than 100 days after setup. Paratone was used as cryoprotectant.The original construct includes a RhoGEF, a PH and a C2 domain. Only the C2 domain was crystallized. The protein preparation probably contains residual E.coli protease that cleaved off the RhoGEF and PH domains and left only C2 domain for crystallization. Crystals of the same cell parameters as that of the structure solved one can also be produced in the presence of 1:100 papain (w/w).
NMR Spectroscopy:
Data Collection:
Data Processing: