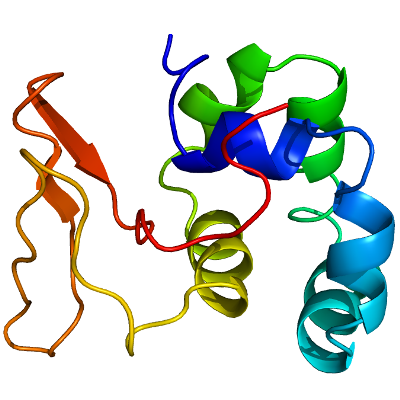
HECW1
PDB:3L4H
Entry Clone Source:Mammalian Gene Collection
SGC Clone Accession:hecw1.AB002320.KZA.ORK01066-KIAA0322.pBluescriptIISKplus
Tag:N-terminal tag: MHHHHHHSSGRENLYFQG
Host:BL-21(DE3)
Vector:pET28a-LIC
Sequence: mhhhhhhssgrenlyfqgSEAESSQSSLDLRREGSLSPVNSQKITLLLQSPAVKFITNPEFFTVLHANYSAYRVFTSSTCLKHMILKVRRDARNFERYQHNRDLVNFINMFADTRLELPRGWEIKTDQQGKSFFVDHNSRATTFIDPRIPLQNG
Growth
Medium:TB
Procedure: Competent BL-21(DE3) cells (Invitrogen, C6000-03) were transformed and grown using the LEX system (HarbingerBiotech) at 37 degC in 2L bottles (VWR, 89000-242) containing 1800 ml of TB (Sigma T0918) supplemented with 150 mM glycerol, 100 µM Kanamycin and 600 µl antifoam 204 (Sigma A-8311). At OD600 = 6, the temperature was reduced to 15 degC, and one hour later the culture was induced with 100 µM IPTG (BioShop IPT001) and incubated overnight (16 hours) at 15 degC. Cell pellets were collected by centrifugation (12,227 xg, 20 mins) and frozen in liquid nitrogen.
Purification
Procedure:
Resin was transferred to a column and washed with 5 column volumes (cv) of Wash buffer A, 5 cv of Wash buffer B, and 5 cv of Wash buffer A. The protein was eluted with 2 cv of Elution buffer. The his-tag was cut with tev (1 mM per 20 mM protein) overnight at 4°C. The protein was further purified by gel filtration through a HighLoad 16/60 Superdex 200 column (GE Healthcare) equilibrated with Gel Filtration buffer. Fractions containing protein (analyzed by ABS280 nm) were pooled and concentrated to 10 mg/ml using concentrators (Amicon) with 5 kDa cutoff. The yield of the protein was approximately 3 mg per L of bacterial culture. Coomassie-stained, SDS-PAGE showed that the product was pure and Mass-spectroscopy by LCMS (Agilent 1100 Series) showed that the protein has only 1 Da more than the calculated molecular weight.
Extraction
Procedure:
Cell pellets were resuspended in Lysis buffer(30 mL per L culture), lysed using a Microfluidizer (Microfluidics, M110-EH) at 18,000 psi, and cleared by centrifugation (40,000 xg for 30 minutes). Cleared lysate was rocked with TALON metal-affinity resin (BD Biosciences) (1.5 mL settled beads per L cell culture) at 4 °C.
Concentration:10.0 mg/ml.
Structure Determination
Crystallization:Crystals were grown at 18 degC using the sitting drop method in 24 well plates (Art Robbins, 102-0004-00) by mixing equal volumes of protein (10.0 mg/ml) (add 0.1 mM Acetic acid pH 3.0 before setting plates) in the presence of trypsin (1 mg trypsin per 500 mg protein) and Crystallization buffer (1.0 M Nacitrate, 0.1mM Immidazole pH 8.0). Suitable crystals were cryoprotected by immersion in well solution supplemented with 15% (v/v) glycerol prior to dunking and storage in liquid nitrogen.
Data Collection:Diffraction data was collected on a crystal of a selenium-methione derivative of the tandem helical box and WW domain of HECW1 at beamline 19-ID at the Argonne Photon Source. The data set was integrated and scaled the HKL2000 program suite.
Data Processing:The structure was solved by single-wavelength anomalous diffraction technique using the program SOLVE. Automated model building using RESOLVE, combined with iterative model building using the graphics program Coot and maximum-likelihood and TLS refinement with the program REFMAC5 led to a model with an Rfactor of 0.15 (Rfree 0.20) for data between 29.45-1.80 Å. Parameters for Translation/liberation/screw (TLS) refinement were generated using the TLSMD web server.