Entry Clone Source: FivePrime |
Entry Clone Accession: TBA - Opher |
SGC Construct ID: STK17BA-c014 |
GenBank GI number: gi|4758194 |
| Vector: pNIC28-Bsa4. Details [PDF ]; Sequence [ FASTA ] or [ GenBank ]
|
Coding DNA sequence:
CATATGCACCATCATCATCATCATTCTTCT
GGTGTAGATCTGGGTACCGAGAACCTGTAC
TTCCAATCCATGGAAAACTTTAATAATTTC
TATATACTTACATCTAAAGAGCTAGGGAGA
GGTAAATTTGCTGTGGTTAGACAATGTATA
TCAAAATCTACTGGCCAAGAATATGCTGCA
AAATTTCTAAAAAAGAGAAGAAGAGGACAG
GATTGTCGGGCAGAAATTTTACACGAGATT
GCTGTGCTTGAATTGGCAAAGTCTTGTCCC
CGTGTTATTAATCTTCATGAGGTCTATGAA
AATACAAGTGAAATCATTTTGATATTGGAA
TATGCTGCAGGTGGAGAAATTTTCAGCCTG
TGTTTACCTGAGTTGGCTGAAATGGTTTCT
GAAAATGATGTTATCAGACTCATTAAACAA
ATACTTGAAGGAGTTTATTATCTACATCAG
AATAACATTGTACACCTTGATTTAAAGCCA
CAGAATATATTACTGAGCAGCATATACCCT
CTCGGGGACATTAAAATAGTAGATTTTGGA
ATGTCTCGAAAAATAGGGCATGCGTGTGAA
CTTCGGGAAATCATGGGAACACCAGAATAT
TTAGCTCCAGAAATCCTGAACTATGATCCC
ATTACCACAGCAACAGATATGTGGAATATT
GGTATAATAGCATATATGTTGTTAACTCAC
ACATCACCATTTGTGGGAGAAGATAATCAA
GAAACATACCTCAATATCTCTCAAGTTAAT
GTAGATTATTCGGAAGAAACTTTTTCATCA
GTTTCACAGCTGGCCACAGACTTTATTCAG
AGCCTTTTAGTAAAAAATCCAGAGAAAAGA
CCAACAGCAGAGATATGCCTTTCTCATTCT
TGGCTACAGCAGTGGGACTTTGAAAACTTG
TTTCACCCTGAAGAAACTTCCAGTTCCTCT
CAAACTCAGGATCATTCTGTAAGGTCCTCT
GAAGACAAGACTTCTAAATCCTCCTGACAG
TAAAGGTGGATACGGATCCGAA |
Final protein sequence:
MHHHHHHSSGVDLGTENLYFQ^SMENFNNF
YILTSKELGRGKFAVVRQCISKSTGQEYAA
KFLKKRRRGQDCRAEILHEIAVLELAKSCP
RVINLHEVYENTSEIILILEYAAGGEIFSL
CLPELAEMVSENDVIRLIKQILEGVYYLHQ
NNIVHLDLKPQNILLSSIYPLGDIKIVDFG
MSRKIGHACELREIMGTPEYLAPEILNYDP
ITTATDMWNIGIIAYMLLTHTSPFVGEDNQ
ETYLNISQVNVDYSEETFSSVSQLATDFIQ
SLLVKNPEKRPTAEICLSHSWLQQWDFENL
FHPEETSSSSQTQDHSVRSSEDKTSKSS
^ TEV cleave site |
Tags and additions: Cleavable N-terminal His6 tag. |
| Host: BL21 (DE3)R3-pRARE2 (Phage resistant strain) |
Growth medium, induction protocol: 5 ml from a 50 ml overnight culture containing 50 µg/ml kanamycin & 34 µg/ml chloramphenicol were used to inoculate each of two 1 litre cultures of LB containing 50 µg/ml kanamycin & 34 µg/ml chloramphenicol. Cultures were grown at 37 °C until the OD600 reached ˜0.5 then the temperature was adjusted to 18 °C. Expression was induced overnight using 0.5 mM IPTG at an OD600 of 0.9. The cells were collected by centrifugation and the pellet re-suspended in binding buffer and frozen.
Binding buffer: 50 mM HEPES pH 7.5; 500 mM NaCl; 5 mM imidazole, 5 % glycerol. |
Extraction buffer, extraction method: Frozen pellets were thawed and fresh 0.5 mM TCEP added to the lysate. Cells were lysed using C5 high pressure homogenizer (Avestin). The lysate was centrifuged at 16,500 rpm for 60 minutes and the supernatant collected for purification. |
Column 1: Ni-affinity. Ni-Sepharose (Amersham), 5 ml of 50 % slurry in 1.5 x 10 cm column, washed with binding buffer. |
Buffers:
Binding buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 5 mM imidazole, 5% Glycerol
Wash buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 25 mM Imidazole, 5% glycerol
Elution buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 50 to 250 mM Imidazole , 5% Glycerol (step elution). |
|
Procedure: The lysate supernatant was loaded by gravity flow on the Ni-sepharose column. The column was then washed with 30 ml wash buffer at gravity flow. The protein was eluted by gravity flow by applying 5-ml portions of elution buffer with increasing concentration of imidazole (50 mM, 100 mM, 150 and 250 mM); fractions were collected until essentially all protein was eluted. |
Column 2: Size Exclusion Chromatography. Superdex S200 16/60 HiLoad |
Buffers: 25 mM HEPES, pH 7.5; 500 mM NaCl, 0.5 mM TCEP |
Procedure: The protein was concentrated and applied to an S200 16/60 HiLoad gel filtration column equilibrated in 25 mM HEPES, pH 7.5; 500 mM NaCl, 0.5 mM TCEP using an ÄKTA express system. |
Mass spectrometry characterization:LC- ESI -MS TOF revealed that the recombinant protein had a mass of 37286 correlating well with the estimated mass of 37285.3 predicted from the sequence of this protein.
|
Protein concentration: Protein was concentrated to 11 mg/ml using an Amicon 10 kDa cut-off concentrator. |
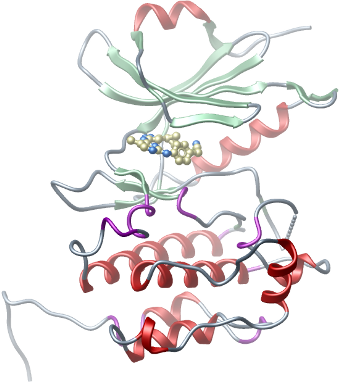
Crystallization: Crystals in complex with ASC68 (K. Shokat, UCSF) were grown at 4 oC in 300 nl sitting drops from a 2:1 ratio of reservoir solution (0.2M (NH4)3(cit)20% PEG 3350) and protein.
Crystals in complex with quercetin (CalBioChem, # : 551600) were grown at 4 °C in 300 nl sitting drops from a 2:1 ratio of protein and reservoir solution containing (0.2M NaCl; 0.1M BIS-TRIS pH 5.5; 25% PEG 3350). |
Data Collection: Crystals were cryo-protected using the well solution supplemented with 25 % Ethylene glycol and flash frozen in liquid nitrogen.
X-ray source:Diffraction data were collected from a single crystal for each dataset on Diamond beamlines IO4 and IO2 at a single wavelength of 0.976 Å and 0.980 Å respectively.
Phasing: The structures were solved by molecular replacement using the structure of human DAPK (PDB ID 3EHA) as a search model. |