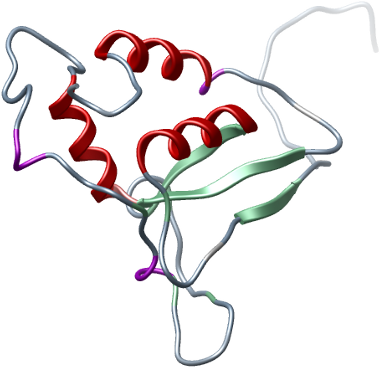
USP15
PDB:3LMN
Entry Clone Accession:BC125123.MGC.CM34-A7.pCR-BluntII-TOPO
Entry Clone Source:Mammalian Gene Collection
SGC Clone Accession:usp15.0001x0133.188G08 (SDC188G08)
Tag:N-terminal tag: MGSSHHHHHHSSGLVPRGS
Host:BL21 (DE3)-V2R
Vector:pET28a-LIC
Sequence:mgsshhhhhhssglvprgsMAEGGAADLDTQRSDIATLLKTSLRKGDTWYLVDSRWFKQWKKYVGFDSWDKYQMGDQNVYPGPIDNSGLLKDGDAQSLKEHLIDELDYILLPTEGWNKLVSWYTLMEGQEPIARKVVEQGMFVKHCKVEVYL
Growth
Medium:TB
Procedure:Competent BL21 (DE3)-V2R cells were transformed and grown using the LEX system (HarbingerBiotech) at 37 °C in 1 L bottles (VWR, 89000-242) containing 800 ml of TB (Sigma, T0918) supplemented with 150 mM glycerol, 100 µM Kanamycin and 600 µl antifoam 204 (Sigma A-8311). When the OD600 reached a value of about 6.0, the temperature was reduced to 15 °C, and one hour later the culture was induced with 100 µM IPTG (BioShop, IPT001) and incubated overnight (16 hours) at 15 °C.
Purification
Procedure: The protein was purified using the Streamline Purification System. Resin was washed with 2x15 ml Washing Buffer and the protein was eluted with 9 ml Elution Buffer. The N-terminal His-tag was removed by overnight incubation of the protein with thrombin (1 U per mg protein) at 4 °C. The buffer was exchanged to Storage Buffer using a desalting column (GE Healthcare, Product Code: 17-0851-01) and the protein sample was concentrated using a 3 KDa MW cut-off concentrator (Millipore, UFC900524) at 3000 x g to a final protein concentration of 25 mg/ml. Coomassie-stained SDS-PAGE showed that the product was pure.
Structure Determination
MassSpec:Mass-spectroscopy by LC/MS (Agilent 1100 Series) showed that the purified protein had the exact molecular weight.
Crystallization:Crystals were grown at 18 degC using the sitting drop method in Intelliplates by mixing equal volumes of protein (15 mg/ml) and Crystallization Buffer (2.0 M sodium formate, 0.1 M sodium acetate, pH 3.8). Suitable crystals were cryoprotected by immersion in well solution supplemented with 20 % (v/v) glycerol prior to dunking and storage in liquid nitrogen.
Data Collection:Diffraction data from a crystal of the DUSP domain of USP15 was collected on a home source Rigaku FR-E, and integrated and scaled using the HKL2000 program suite.
Data Processing:The structure was solved by molecular replacement techniques using the program PHASER and search model PDB entry 3JYU. Automated model building using ARP/wARP, combined with iterative model building using the graphics program Coot and maximum-likelihood and TLS refinement with the program REFMAC5 led to a model with an R factor of 18.3% (Rfree 20.4%) for data between 19.5-2.15 Å. Parameters for Translation/liberation/screw (TLS) refinement were generated using the TLSMD web server.