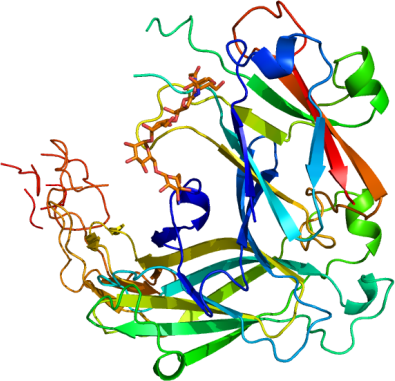
EPHA2 EPHA1
PDB:3MBW
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:epha2.BC037166.MGC.AU80A3.pOTB7
efna1a.BC032698.MGC.AT54G10.pCMVSPORT6
Entry Clone Source:MGC
SGC Clone Accession:epha2.023.326.121A07 (SDC121A07)
efna1a.017.171.121F08 (SDC121F08)
Tag:N-terminal tag: APEHHHHHHDYDIPTTENLYFQGAMD
Host:DH10Bac E.coli cells (Invitrogen, 10361-012)
Construct
Prelude:The Ligand Binding Domain (LBD) and Cysteine-rich Domain (CRD) of Ephrin receptor A2 (EphA2, also known as Eck) was cloned from a cDNA template from the Mammalian Gene Collection (epha2.BC037166.MGC.AU80A3.pOTB7) into the pFHMSP-LIC-N vector (SGC, ) using the In-Fusion CF Dry-Down PCR Cloning Kit (Clontech, 639605), resulting in a plasmid called epha2.023.326.121A07 (SDC121A07). The globular domain of Ephrin-A1 (EFNA1, also known as B61, EFL1, ECKLG, EPLG1, LERK1, and TNFAIP4) was cloned from a cDNA template from the Mammalian Gene Collection (efna1a.BC032698.MGC.AT54G10.pCMVSPORT6) into the pFHMSP-LIC-N vector (SGC) using the In-Fusion CF Dry-Down PCR Cloning Kit (Clontech, 639605), resulting in a plasmid called efna1a.017.171.121F08 (SDC121F08).
Sequence: Construct epha2.023.326.121A07: apehhhhhhdydipttenlyfqgamdAAQGKEVVLLDFAAAGGELGWLTHPYGKGWDLMQNIMNDMPIYMYSVCNVMSGDQDNWLRTNWVYRGEAERIFIELKFTVRDCNSFPGGASSCKETFNLYYAESDLDYGTNFQKRLFTKIDTIAPDEITVSSDFEARHVKLNVEERSVGPLTRKGFYLAFQDIGACVALLSVRVYYKKCPELLQGLAHFPETIAGSDAPSLATVAGTCVDHAVVPPGGEEPRMHCAVDGEWLVPIGQCLCQAGYEKVEDACQACSPGFFKFEASESPCLECPEHTLPSPEGATSCECEEGFFRAPQDPASMPCT Construct efna1a.017.171.121F08:apehhhhhhdydipttenlyfqgamdAADRHTVFWNSSNPKFRNEDYTIHVQLNDYVDIICPHYEDHSVADAAMEQYILYLVEHEEYQLCQPQSKDQVRWQCNRPSAKHGPEKLSEKFQRFTPFTLGKEFKEGHSYYYISKPIHQHEDRCLRLKVTVSGKITHSPQAHVNPQEKRLAADDP
Vector:pFHMSP-LIC-N vector (SGC)
Growth
Medium:Serum Free Medium (HyClone SFX-Insect, SH3027802)
Antibiotics:
Procedure:These plasmids were transformed into DH10Bac E.coli cells (Invitrogen, 10361-012) and a mini-prep was performed to obtain recombinant viral bacmid DNA. SF9 cells (Invitrogen, 11496-015) were transfected with bacmid using Cellfectin reagent (Invitrogen, cat # 10362-010), and recombinant baculovirus was generated. Viral stock was amplified from P1 to P3.
Sf9 cells grown in Serum Free Medium (HyClone SFX-Insect, SH3027802) at density of 3.5 million cells per milliliter of media and with viability not less then 97 % were infected with 5 mL of P3 viral stock for each 1 L of cell culture. Cell culture media was collected after 3-4 days of incubation on a shaker at 100 RPM and 27 °C when cells viability dropped to 45-65 %.
Purification
Procedure
A 3.2 L volume of medium was mixed with 30 ml pre-equilibrated NiNTA Superflow beads (Qiagen, 30450) and stirred (Talboys/Troemner) for 1 hour on ice. The resin was transferred to a 100 ml column, washed with 300 ml of 50 mM Tris pH 8.0, 500 mM NaCL, 5mM imidazole, and the protein was eluted with 30-50 ml of 50 mM Tris pH 8.0, 500 mM NaCL, 300 mM imidazole, 5% glycerol. A second round of NiNTA batch absorption may have been performed for increased yield. The eluate was dialyzed against 50-100 X volume of 50 mM Tris pH 8.0 150 mM NaCL overnight at 4 degC. Purified protein was concentrated using 15 mL concentrators with an appropriate molecular weight cut-off (Amicon Ultra-15 10,000 MWCO, Millipore, UFC 901024) to a final value of 10 mg/mL. Average yield was about 2 and 10 mg/L for epha2.023.326 and efna1a.017.171, respectively. Coomassie-stained SDS-PAGE showed that the products were pure and Mass-spectroscopy by LCMS showed that the proteins were glycosylated.
Extraction
Procedure
Cultured media was centrifuged at 14,000 xg for 15 minutes and the pH of the supernatant was adjusted to 7.5 at room temperature by adding 500 mM Tris pH 8.0, 1500 mM NaCL. Protease inhibitors were added to final concentrations of 1 mM (phenylmethanesulfonyl fluoride, PMSF, Bioshop) and 2 mM (benzamidine hydrochloride, Sigma).
Concentration:10 mg/mL.
Ligand
MassSpec:
Crystallization:Mixture of EphA2 (at 10 g/L) and efna1a (at 22 g/L) in ratio 0.02 uM : 0.02 uM were set (500 nL protein + 500 nL well solution) at 18 ºC. The crystal that was used to collect data was grown using the sitting drop method in IntelliPlates 96-2 well (Hampton Research, HR3-299) in Crystallization Buffer (10 % PEG 3350, 0.16 M NH4PO4). Suitable crystals were cryoprotected by immersion in 1 uL CB and 1 uL cryoprotectant (20% (w/v) sucrose, 4% (w/v) glucose, 18% (v/v) glycerol and 18% (v/v) ethylene glycol prior to dunking and storage in liquid nitrogen.
NMR Spectroscopy:
Data Collection:Diffraction data from a crystal of EphA2 (residues 23-326) in complex with EphA1 (residues 17-171) was collected at beamline 19-ID at the Argonne Photon Source. The data set was integrated and scaled using the HKL2000 program suite.
Data Processing:The structure was solved by molecular replacement techniques using the program PHASER and search model PDB entries 3C8X and 1SHW. Iterative model building using the graphics program Coot and maximum-likelihood and TLS refinement with the programs REFMAC5 and BUSTER led to a model with an R factor of 22.6% (Rfree 27.2%) for data between 34.1-2.81Å. Parameters for Translation/liberation/screw (TLS) refinement were generated using the TLSMD web server. Statistics of data collection, processing, and refinement are provided in Figure 3. The coordinates and structure factors have been deposited on 2010.03.26 into the RCSB PDB database with ID code 3MBW.