Entry Clone Source: Site directed mutagenesis |
Entry Clone Accession: n/a |
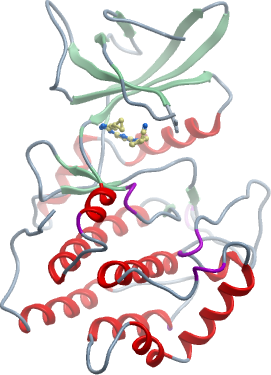
SGC Construct ID: VRK1A-c510 |
GenBank GI number: gi|4507903 |
Vector: pNIC28-Bsa4. Details [PDF ]; Sequence [ FASTA ] or [ GenBank ] |
Amplified construct sequence:
Nucleotides in bold and underlined were multated from refseq in an attempt to reduce surface entropy in the translated protein.
CATATGCACCATCATCATCATCATTC
TTCTGGTGTAGATCTGGGTACCGAGA
ACCTGTACTTCCAATCCATGCGTGTA
AAAGCAGCTCAAGCTGGAAGACAGAG
CTCTGCAAAGAGACATCTTGCAGAAC
AATTTGCAGTTGGAGAGATAATAACT
GACATGGCAAAAAAGGAATGGAAAGT
AGGATTACCCATTGGCCAAGGAGGCT
TTGGCTGTATATATCTTGCTGATATG
AATTCTTCAGAGTCAGTTGGCAGTGA
TGCACCTTGTGTTGTAAAAGTGGAAC
CCAGTGACAATGGACCTCTTTTTACT
GAATTAAAGTTCTACCAACGAGCTGC
AAAACCAGAGCAAATTCAGAAATGGA
TTCGTACCCGTAAGCTGAAGTACCTG
GGTGTTCCTAAGTATTGGGGGTCTGG
TCTACATGACAAAAATGGAAAAAGTT
ACAGGTTTATGATAATGGATCGCTTT
GGGAGTGACCTTCAGAAAATATATGA
AGCAAATGCCAAAAGGTTTTCTCGGA
AAACTGTCTTGCAGCTAAGCTTAAGA
ATTCTGGATATTCTGGAATATATTCA
CGAGCATGAGTATGTGCATGGAGATA
TCAAGGCCTCAAATCTTCTTCTGAAC
TACAAGAATCCTGACCAGGTGTACTT
GGTAGATTATGGCCTTGCTTATCGGT
ACTGCCCAGAAGGAGTTCATAAAGAA
TACAAAGAAGACCCCAAAAGATGTCA
CGATGGCACTATTGAATTCACGAGCA
TCGATGCACACAATGGCGTGGCCCCA
TCAAGACGTGGTGATTTGGAAATACT
TGGTTATTGCATGATCCAATGGCTTA
CTGGCCATCTTCCTTGGGAGGATAAT
TTGAAAGATCCTAAATATGTTAGAGA
TTCCAAAATTAGATACAGAGAAAATA
TTGCAAGTTTGATGGACAAATGTTTT
CCTGAGAAAAACAAACCAGGTGAAAT
TGCCAAATACATGGAAACAGTGAAAT
TACTAGACTACACTGAAAAACCTCTT
TATGAAAATTTACGTGACATTCTTTT
GCAAGGACTAAAAGCTATAGGAAGTA
AGGATGATGGCAAATTGGACCTCAGT
GTTGTGGAGAATGGAGGTTTGAAAGC
AAAAACAATAACAAAGAAGCGAAAGA
AAGAAATTGAAGAATGACAGTAAAGG
TGGATACGGATCCGAA
|
Final protein sequence (Tag sequence in lowercase):
mhhhhhhssgvdlgtenlyfqs^MMH
HHHHHSSGVDLGTENLYFQSMRVKAA
QAGRQSSAKRHLAEQFAVGEIITDMA
AAAWKVGLPIGQGGFGCIYLADMNSS
ESVGSDAPCVVKVEPSDNGPLFTELK
FYQRAAKPEQIQKWIRTRKLKYLGVP
KYWGSGLHDKNGKSYRFMIMDRFGSD
LQKIYEANAKRFSRKTVLQLSLRILD
ILEYIHEHEYVHGDIKASNLLLNYKN
PDQVYLVDYGLAYRYCPEGVHKAYAA
DPKRCHDGTIEFTSIDAHNGVAPSRR
GDLEILGYCMIQWLTGHLPWEDNLKD
PKYVRDSKIRYRENIASLMDKCFPAA
NAPGEIAKYMETVKLLDYTEKPLYEN
LRDILLQGLKAIGSKDDGKLDLSVVE
NGGLKAKTITKKRAAEIEE
^ TEV cleave site |
Tags and additions: Cleavable N-terminal His6 tag. |
Host: BL21 (DE3)R3-pRARE2 (Phage resistant strain).
|
Growth medium, induction protocol: The expression plasmid was transformed into the host strain and plated on LB-agar containing 50µg/ml kanamycin and 35µg/ml chloramphenicol. Several colonies were combined to inoculate a 1ml culture in TB (+ 50µg/ml kanamycin, 35µg/ml chloramphenicol). The culture was grown overnight, glycerol was added to 15% v/v (from a 60% stock), and the resulting glycerol stock was frozen at -80°C in 100 µl aliquots. A loopful of cells from the glycerol stock was inoculated into 5ml of LB medium containing 100µg/ml kanamycin and 35µg/ml chloramphenicol and grown overnight at 37°C. Overnight culture was used to inoculate 1 liter of LB containing 50µg/ml kanamycin & 34µg/ml chloramphenicol. Culture was grown at 37°C until the OD600 reached ~3 and then cooled to 18°C for 1 hour. IPTG was added to 0.1 mM, and growth continued at 18°C overnight. The cells were collected by centrifugation then pellet re-suspended in 2x lysis buffer and frozen.
Lysis buffer: 50 mM HEPES buffer, pH 7.5, 500 mM NaCl, 10 mM imidazole, 0.5 mM TCEP, 1x Protease Inhibitors Cocktail Set VII (Calbiochem, 1/1000 dilution), and 15 units/ml Benzonase. 2x Lysis buffer contains the same components at double concentration.
Extraction buffer, extraction method: Frozen pellets were thawed and fresh 0.5 mM TCEP added to the lysate. Cells were lysed by high pressure homogenization (20 kpsi). The lysate was centrifuged at 16,500 rpm for 60 minutes and the supernatant collected for purification. |
Column 1: Ni-affinity, HisTrap Crude FF, 5 ml (GE Healthcare). |
Column 1 Buffers:
Affinity buffer: 50 mM HEPES buffer, pH 7.5, 500 mM NaCl, 10 mM imidazole, 5% Glycerol, 0.5 mM TCEP.
Wash buffer: 50 mM HEPES buffer, pH 7.5, 500 mM NaCl, 30 mM imidazole, 5% Glycerol, 0.5 mM TCEP.
Elution buffer: 50 mM HEPES buffer, pH 7.5, 500 mM NaCl, 300 mM imidazole, 5% Glycerol, 0.5 mM TCEP. |
Column 1 Procedure: The cell extract was loaded on the column at 4 ml/minute on an ÄKTA-express system (GE Healthcare). The column was washed with 10 volumes of lysis buffer, 10 volumes of wash buffer, and then eluted with elution buffer at 4 ml/min. The eluted peak of A280 was automatically collected. |
Column 2: Gel filtration, Hiload 16/60 Superdex S75 prep grade, 120 ml (GE Healthcare) |
Column 2 Buffer: 50 mM HEPES, pH 7.5, 500 mM NaCl, 5% Glycerol, 0.5 mM TCEP. |
Column 2 Procedure: The eluted fractions from the Ni-affinity Histrap column were loaded on the gel filtration column in GF buffer at 1.2 ml/min. Eluted proteins were collected in 2-ml fractions and analyzed on SDS-PAGE. |
TEV protease digestion: Peak fractions containing VRK1 were pooled and TEV protease was added at a molar ratio of 1:30. The digestion was left overnight at 4°C. His-TEV and contaminating proteins were removed by passing to 300µl Ni resin pre-equilibrated in GF buffer. The flow through containing TEV-cleaved protein was collected, concentrated to 30mg/ml using a centricon centrifugal device with a 30kDa MWCO, frozen at -80°C in 500µl aliquots. |
Mass spectrometry characterization: Measured: 41078, 41163 (one phosphorylation site) and 41240 (2 phosphorylation sites). Expected mass of unmodified protein: 41076.2. |
Protein concentration: 2.5 mg of VRK1 was diluted into 15 ml of low salt buffer (5mM HEPES. pH 7.5, 200 mM NaCl, 1% glycerol). 2 molar excess of ligand was added to diluted VRK1 and protein-ligand complex was concentrated to 20 mg/ml using an Amicon 30 kDa cut-off concentrator. |
Crystallisation: Prior to crystallization, pure protein at 20 mg/ml was incubated with 1mM K00604a (ASC24, provided by the Shokat laboratory) compound. Optimized condition contained 0.2 M K/Na(tart) and 20% PEG 3350, pH 7.0. Viable crystals were obtained at 4°C in a sitting drop mixing protein and reservoir solution at 2:1 volume ratio. |
Data Collection: The crystals were cryo-protected with the mother liquor containing 25% ethylene glycol and flash frozen in liquid nitrogen. Diffraction data were collected at Diamond Beamline I02 using monochromatic radiation at wavelength 0.9790 Å.
Phasing: Structure of VRK1 was solved by molecular replacement method using an ensemble of the following PDB codes as a search model; 2V62, 2JII, 1CKJ. |