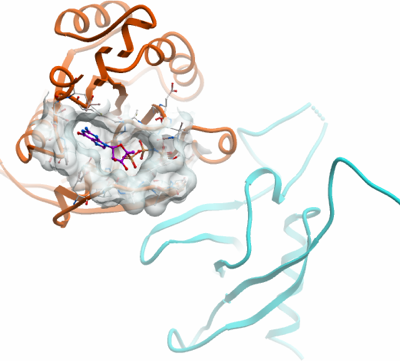
PLXNA2 + RND1
PDB:3Q3J
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:PLXNA2: SGC cDNA library: 20-B7
Rnd1: MGC cDNA library: AT22-F2
Entry Clone Source:PLXNA2: SGC cDNA library
Rnd1: MGC cDNA library
SGC Clone Accession:PLXNA2: HPC09R-B07
Rnd1: HPC060-H10
Tag:mhhhhhhssgrenlyfq*g
Host:BL21-V2R-pRARE2
Construct
Prelude:PLXNA2:L1490-K1600
Tag was removed
Rnd1:R5-V200
Tag was not removed
Sequence:PLXNA2:gLIRQQIEYKTLILNCVNPDNENSPEIPVKVLNCDTITQVKEKILDAVYKNVPYSQRPRAVDMDLEWRQGRIARVVLQDEDITTKIEGDWKRLNTLMHYQVSDRSVVALVPK
RND1:mhhhhhhssgrenlyfqgRAPQPVVARCKLVLVGDVQCGKTAMLQVLAKDCYPETYVPTVFENYTACLETEEQRVELSLWDTSGSPYYDNVRPLCYSDSDAVLLCFDISRPETVDSALKKWRTEILDYCPSTRVLLIGCKTDLRTDLSTLMELSHQKQAPISYEQGCAIAKQLGAEIYLEGSAFTSEKSIHSIFRTASMLCLNKPSPLPQKSPV
Vector:pET28-mhl (GI:134105571)
Growth
Medium:
Antibiotics:
Procedure:LEX Bubbling. The target protein was expressed in E. coli by inoculating 100 mL of overnight culture grown in Luria-Bertani medium into a 2 L of Terrific Broth medium in the presence of 50 µg/mL kanamycin and 25 µg/mL chloramphenicol at 37 °reeC. When OD600 reached ~3.0, the temperature of the medium was lowered to 15 °reeC and the culture was induced with 1 mM IPTG. The cells were allowed to grow overnight before harvested and flash frozen in liquid nitrogen and stored at -80 °reeC.
Purification
Procedure
The lysate was centrifuged at 16,000 rpm for 1 hour and the supernatants were mixed with 6 mL 50% flurry of Ni-NTA beads and incubated at 4°reeC on rotary shaker for 1 hour. The mixture was then centrifuged at 2300 rpm for 5 min and the supernatant wasdiscarded. The beads were then washed with washing buffer and finally the elution buffer. The flow-trough was collected and further purified by a Superdex-75 gel filtraton column pre-equilibrated with gel filtration buffer.
For PLXNA2, the protein sample was treated with TEV during overnight dialysis at room temperature before loading to the gel filtration column, and the excesses TEV was removed by adding 100 uL Co-NTA beads, the followthrough was collected for further gel filtration purification.
Fractions containing the protein were collected and concentrated with Amicon Ultra-15 centrifugal filter (M.W.cut-off 3,000). The purity of the preparation is tested by SDS-PAGE to be greater than 95%.
During purification, the tag of RND1 was not removed.
Gel filtration buffer for the complex: 20 mM Tris pH 6.5, 500 mM NaCl, 5% Glycerol, 10mM TCEP, 5mM MgCl2
PLXNA2 and RND1 were mixed at a molar ratio of 1:1 with 5mM GNP, and incubated at 4 degree for overnight. Fractions containing the complex were collected in further gel filtration and concentrated with Amicon Ultra-15 centrifugal filter (M.W.cut-off 3,000).
Extraction
Procedure
Frozen cells from 2L TB culture were thawed and resuspended in 250 mL extraction buffer with freshly added 0.5% CHAPS, and supplemented with 1.7 &milliL protease inhibitor cocktail (SIGMA Catalog # P8849), and 10 µL benzonase (Sigma Catalog # E1014, 250U/µL), and lysed using sonication at 120W for 6 minutes on 50% duty cycle.
Concentration:PLXNA2:14.6 mg/mL
Rnd1: 6.0 mg/mL
PLXNA2-RND1 complex: 2.7 mg/mL
Ligand
GNPMassSpec:PLXNA2: expected 12990.96, measured 12991.46
RND1: expected 24060.48, measured 24060.58
Crystallization:The complex sample for crystallization was incubated with 5mM GNP. Crystallization was setup in sitting drops with Red Wings and SGC-I screens initially. After 5 months, the crystals were found in 25.0% P3350, 0.2 M MgCl2, 0.1 M HEPES buffer pH 7.5. The optimization has been carried on with the initial condition in hanging drops. Crystals were reproduced in one week. Diffracting crystals were from further seeding in sitting drops with the seeds of the optimization crystals.
Crystal used for structure determination was grown in 25.5% P3350, 0.2 M MgCl2, 0.1 M HEPES buffer pH 7.5, with the seed dilution ratio of 1:10000 in sitting drop setup. Crystals grow to a mountable size after 2 weeks.
Paratone was used as cryoprotectant.
NMR Spectroscopy:
Data Collection:
Data Processing: