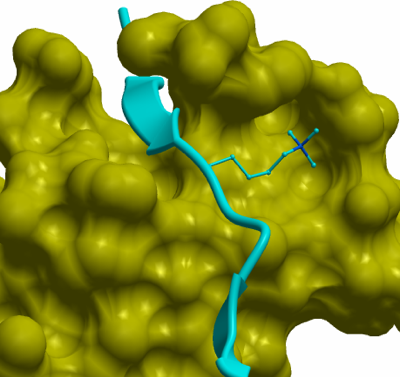
The human MPP8 chromo domain adopts a canonical chromo domain fold similar to that of the chromo domains of HP1 and Polycomb and it consists of 3 beta strands and one long helix at C-terminus. In the structure of the hMPP8 chromodomain in complex with the H3K9me3 peptide, two histone H3K9me3 peptides bind to the opposite faces of the hMPP8 chromodomain homodimer, respectively. Structural comparison of the hMPP8 chromodomain-H3K9me3 peptide complex and the free hMPP8 chromodomain identified a newly formed β strand (named β1) by the N-terminal residues, which exited as a loop in the free hMPP8 chromodomain structure. This β strand is induced by the contact with the H3 tail peptide, which was observed in the structures of Drosophila HP1 and Polycomb chromodomain in complex with methyllysine histone peptides before.
From the complex structure we can see that the H3K9me3 peptide binds to hMPP8 chromodomain in a cleft between the N-terminal newly formed β1 strand and the loop connecting β4 and αA. Similar to the structure of the Drosophila HP1 and Polycomb chromodomain in complex with methyllysine histone peptides, the interactions between hMPP8 chromodomain and H3K9me3 largely involve the main chains of both the protein and the peptide, including the residues Gln5, Thr6, Ala7, and Arg8 of the H3 tail and the residues Val58, Phe59, Glu60, and Val61 located at the β1 strand in hMPP8 chromodomain. In addition, the residues of Gln5 and Arg8 form van der waals contacts with the residues of 98-100 located in the loop connecting β4 and αA, whereas Gln5 and Ser10 form hydrogen-bonds with residues of Glu101, Val102 and Glu91, respectively. As demonstrated in most complex structures of methyllysine peptides and their recognition modules, the trimethylated K9 lies in a hydrophobic pocket formed by three aromatic residues, Phe59, Trp80, and Tyr83. And the trimethyl-K9 is anchored by cation-π and van der Waals interactions within this aromatic cage.