Entry Clone Source: MGC |
Entry Clone Accession: IMAGE:5181100 |
SGC Construct ID: AMPHA-c013 |
GenBank GI number: gi|4502081 |
Vector: pNIC28-Bsa4. Details [PDF ]; Sequence [ FASTA ] or [ GenBank ] |
Amplified construct sequence:
ATGCACCATCATCATCATCATTCTTC
TGGTGTAGATCTGGGTACCGAGAACC
TGTACTTCCAATCCATGACAAAAGAC
GAACAGTTCGAAGAATATGTCCAGAA
CTTCAAACGGCAAGAAGCAGAGGGTA
CCAGACTTCAGCGAGAACTCCGAGGA
TATTTAGCAGCAATCAAAGGCATGCA
GGAGGCCTCCATGAAGCTCACAGAGT
CGCTGCATGAAGTCTATGAGCCTGAC
TGGTATGGGCGGGAAGATGTGAAAAT
GGTTGGTGAGAAATGTGATGTGCTGT
GGGAAGACTTCCATCAAAAACTCGTG
GATGGGTCCTTGCTAACACTGGATAC
CTACCTGGGGCAATTTCCTGACATAA
AGAATCGCATCGCCAAGCGCAGCAGG
AAGCTAGTGGACTATGACAGTGCCCG
CCACCATCTGGAAGCTCTGCAGAGCT
CCAAGAGGAAGGATGAGAGTCGAATC
TCTAAGGCAGAAGAAGAATTTCAGAA
AGCACAGAAAGTGTTTGAAGAGTTTA
ACGTTGACTTACAAGAAGAGTTACCA
TCATTATGGTCAAGACGAGTTGGATT
TTATGTTAATACTTTCAAAAACGTCT
CCAGCCTTGAAGCCAAGTTTCATAAG
GAAATTGCGGTGCTTTGCCACAAACT
GTATGAAGTGATGACAAAACTGGGTG
ACTGA
|
Final protein sequence (Tag sequence in lowercase):
mhhhhhhssgvdlgtenlyfq^smTK
DEQFEEYVQNFKRQEAEGTRLQRELR
GYLAAIKGMQEASMKLTESLHEVYEP
DWYGREDVKMVGEKCDVLWEDFHQKL
VDGSLLTLDTYLGQFPDIKNRIAKRS
RKLVDYDSARHHLEALQSSKRKDESR
ISKAEEEFQKAQKVFEEFNVDLQEEL
PSLWSRRVGFYVNTFKNVSSLEAKFH
KEIAVLCHKLYEVMTKLGD
^ TEV cleavage site |
Tags and additions: Cleavable N-terminal His6 tag. |
Host: BL21 (DE3)R3-pRARE2 (Phage resistant strain).
|
Growth medium, induction protocol: The expression plasmid was transformed into the host strain and plated on LB-agar containing 50µg/ml kanamycin and 34µg/ml chloramphenicol. Several colonies were combined to inoculate a 1ml culture in TB (+ 50µg/ml kanamycin, 34µg/ml chloramphenicol). The culture was grown overnight, glycerol was added to 15% v/v (from a 60% stock), and the resulting glycerol stock was frozen at -80°C in 100µl aliquots. A loopful of cells from the glycerol stock was inoculated into 20ml of TB medium containing 50µg/ml kanamycin and 34µg/ml chloramphenicol and grown overnight at 37°C. 2x1L TB medium (containing 50µg/ml kanamycin) were each inoculated with 10ml of the overnight culture and grown in 2.5L UltraYield baffled flasks until OD600 of 3.0. Cells were cooled to 18°C, IPTG added to 0.1 mM and growth continued at 18°C overnight. The cells were collected by centrifugation then the pellets were scraped out and transferred to 50ml Falcon tubes and frozen at -80°C.
Lysis buffer: 25 mM Tris buffer, pH 8.0; 500 mM NaCl; 5 mM Imidazole; 10% glycerol; 0.5 mM TCEP; 1x Protease Inhibitors Cocktail Set VII (Calbiochem, 1/1000 dilution).
2x Lysis buffer contains the same components at double concentration.
Extraction method: Frozen cell pellets (86g) were thawed briefly in a bath of warm water (20 - 37°C) then transferred to ice. One volume (i.e. 1ml for every gram of cells) of 2x lysis buffer was added, followed by 1x lysis buffer to a total volume of 300ml. The cells were resuspended by agitating and disrupted by sonication. Nucleic acids and cell debris were removed by adding 0.15% PEI (polyethyleneimine) from a 5% (w/v, pH 7.5) stock, stirring for 15 minutes, then centrifugation for 1 hour at 25,000g. |
Column 1: Histrap FF 5ml (GE Healthcare). |
Column 1 Buffers:
Affinity buffer: 25 mM Tris, pH 8.0; 100 mM KCl; 10% glycerol; 0.5 mM TCEP; 30 mM Imidazole.
Wash buffer: 25 mM Tris, pH 8.0; 100 mM KCl; 10% glycerol; 0.5 mM TCEP; 30 mM Imidazole.
Elution buffer: 25 mM Tris, pH 8.0; 100 mM KCl; 10% glycerol; 0.5 mM TCEP; 300 mM Imidazole. |
Column 1 Procedure: The cell extract was loaded onto the column at 5ml/min on an ÄKTA-express system (GE Healthcare). The column was then washed with 10 volumes of loading buffer, 10 volumes of wash buffer, then eluted with elution buffer at 4ml/min. The eluted peak of A280 was automatically collected. |
Enzymatic treatment: The eluted protein was treated with TEV protease (His6-tagged) overnight at 4°C, while being dialysed in 25 mM Tris, pH 8, 100 mM KCl, 10% glycerol, 1 mM TCEP, using 3000 MWCO dialysis tubing. The protease and other impurities were removed by passing through a 1ml Histrap column. |
Column 2: Gel filtration, HiLoad 16/60 Superdex 75, 120ml. |
GF Buffer: 25 mM Tris, pH 8.0; 100 mM KCl; 10% glycerol; 1 mM TCEP. |
Column 2 Procedure: The protein was loaded and fractionated on the gel filtration column in GF buffer at 1.2ml/min. 2ml fractions were collected at the A280 peaks. The fractions were analyzed by SDS-PAGE and relevant fractions were pooled. This column resulted in a unique peak of the AMPH protein, at an elution volume compatible with a dimer.
|
Protein concentration: The protein was concentrated to 14.5mg/ml (concentrations estimated by A280, using an extinction coefficient of 28480 and molecular mass of 24180Da). |
Mass spectrometry characterization: ESI-TOF Mass spectrometry revealed a mass of 24,181.41Da, compared to the predicted 24,180.4Da (after tag cleavage). |
Crystallisation: Crystals were grown by vapor diffusion at 4°C in 150nl sitting drops. A crystal was grown by mixing 75nl of protein solution (14.5mg/ml) and 75nl of precipitant consisting of 2.66% PEG 6000, 2.66% PEG 8000, 2.66% PEG 10,000, 0.02 M Magnesium Acetate, 0.05 M Potassium Chloride 0.1 M MES pH 6.2. Crystals appeared in about a week at 4°C. Crystals were cryo-protected by bathing the crystal in mother liquor supplemented with 25% Ethylene glycol and flash-freezing in liquid nitrogen. This crystal was used to collect a native dataset.
Another crystal was grown by mixing 50nl of protein solution and 100nl of precipitant consisting of 2.66% PEG 6000, 2.66% PEG 8000, 2.66% PEG 10,000, 0.05 M Magnesium Acetate, 0.02 M Potassium Chloride 0.1 M MES pH 6.2. Crystals appeared in about a week at 4°C. The crystals were soaked for 35 minutes with 5 mM Thiomersal, before cryoprotection in mother liquor supplemented with 25% Ethylene glycol and flash-freezing in liquid nitrogen.
|
Data collection: Diffraction data for the native dataset were collected from a single crystal at the Diamond synchrotron beamline I04-1, at a wavelength of 0.9795Å.
Diffraction data for the derivatised crystal was collected from a single crystal on a Bruker in-house source.
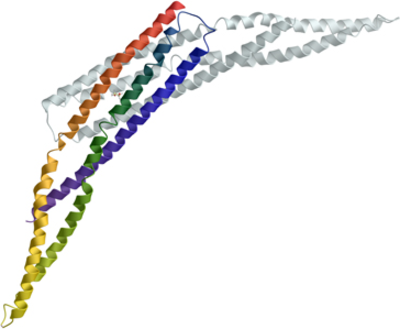
The protein crystallised in space group P31 1 2 with one molecule in the asymmetric unit. Initial phase estimations using molecular replacement of the native resolution data set proved insufficient to build and refine the model. 2 mercury sites were unambiguously identified using SHELXD, with an anomalous signal extending to 4-5Å. The derivatised crystal dataset was used with the native dataset in a SIRAS experiment in SHARP, with solvent flattening in SOLOMAN, resulting in an electron density map clearly showing distinctive coiled-coils, typical of BAR domains. The model was built in COOT, utilising the heavy-atom sites to guide placement of the amino acid sequence register, and refined to a final resolution of 2.3Å using BUSTER-TNT.
|