Molecular Biology
Entry Clone Accession: n/a
Entry Clone Source: Synthetic
SGC Construct ID: CREBBPA-c003
Protein Region: R1081-G1197
Vector: pNIC28-Bsa4
Tag: N-6HIS;N-TEV
Host: BL21(DE3)-R3-pRARE2
Sequence (with tag(s)): MHHHHHHSSGVDLGTENLYFQSMRKKIFKPEELRQALMPTLEALYRQDPESLPFRQPVDPQLLGIPDYFDIVKNPMDLSTIKRKLDTGQYQEPWQYVDDVWLMFNNAWLYNRKTSRVYKFCSKLAEVFEQEIDPVMQSLG
Sequence after tag cleavage: SMRKKIFKPEELRQALMPTLEALYRQDPESLPFRQPVDPQLLGIPDYFDIVKNPMDLSTIKRKLDTGQYQEPWQYVDDVWLMFNNAWLYNRKTSRVYKFCSKLAEVFEQEIDPVMQSLG
DNA Sequence: CATATGCACCATCATCATCATCATTCTTCTGGTGTAGATCTGGGTACCGAGAACCTGTACTTCCAATCCATGCGCAAAAAAATTTTTAAACCGGAAGAACTGCGTCAGGCGCTGATGCCGACCCTGGAAGCGCTGTATCGCCAGGATCCGGAAAGCCTGCCGTTTCGTCAGCCGGTGGATCCGCAGCTGCTGGGTATCCCGGATTATTTTGATATTGTGAAAAACCCGATGGATCTGAGCACCATCAAACGCAAACTGGATACCGGCCAGTATCAGGAACCGTGGCAGTATGTGGATGATGTTTGGCTGATGTTTAATAATGCGTGGCTGTATAACCGTAAAACCAGCCGTGTGTATAAATTCTGTAGCAAACTGGCGGAAGTTTTTGAACAGGAAATTGATCCGGTGATGCAGAGCCTGGGCTGACAGTAAAGGTGGATACGGATCCGAA
Protein Expression
Medium: Terrific Broth
Antibiotics: Kanamycin
Procedure: Host cells transformed with the expression plasmids were plated out onto LB-agar plates containing 50 μg/ml kanamycin. The next day several colonies were combined into 1 ml TB (Terrific Broth), 50 μg/ml kanamycin, which was then grown overnight and stored as glycerol stocks at -80°C. The glycerol stock was used to inoculate a 10-ml starter culture in TB + kanamycin (50 μg/ml). This starter culture was grown overnight at 37°C and used to inoculate a 1 liter culture in the same medium. The culture was grown in baffled flasks at 37°C until the OD600 reached ~3.5. After that the temperature was lowered to 18°C. Protein production was induced with 0.1mM IPTG and the recombinant bromodomain was incubation continued at 18C overnight. The next day cells were harvested by centrifugation at 4000 rpm for 30 minutes. The cell pellet was stored at -80°C degrees
Protein Purification
Procedure: Α cell pellet (40g) from 4 L cultue was re-suspended in one volume (40 ml) of lysis buffer. The re-suspended cells were lysed by one passage through a Constant Systems cell breaker and subsequent sonication; the cell breaker was washed with 1x extraction buffer, bringing the total volume to 120 ml. DNA was precipitation by addition of polyethyleneimine (PEI, pH 7.5) to a final concentration of 0.15 % during an incubation time of 30 min on ice, followed by a centrifugation at 17,000 rpm (4°C); The supernatant was further cleared by filtration through a 0.2 µm serum Acrodisc filter. Fractions containing the expressed bromo domain were collected after his6-tag cleavage and were loaded on a SEC column at 1.0 ml/min. Eluted fractions were >95% pure as judged by SDS-PAGE
Buffers:
Lysis buffer (500mM NaCl, 50mM pH8.0 KH2PO4, 0.5mM TCEP, Benzonase 1μl/15 ml buffer, Protease inhibitor (1 μl/ml))
Affinity binding buffer: 10mM Imidazole, 500mM NaCl, 50mM pH8.0 KH2PO4, 0.5mM TCEP
Columns:
Column 1: Ni-affinity chromatography: HisTrap FF Crude, 5 ml (GE Healthcare).
Binding Buffer: 50mM NaH2PO4, 500mM NaCl, 30mM Imidazole, pH 8.0, 0.5mM TCEP.
Elution Buffer: 50mM NaH2PO4, 500mM NaCl, 250mM Imidazole, pH 8.0, 0.5mM TCEP.
Column 2: Size exclusion chromatography HiLoad 16/60 Superdex 75
SEC-Buffers: 10mM Hepes, pH 7.4, 500 mM NaCl, 5% glycerol, 0.5mM TCEP
Concentration: 10.6 mg/ml
Mass-spec Verification: 14208.6004
Compound Exact Mass: 261.100108
Structure Determination
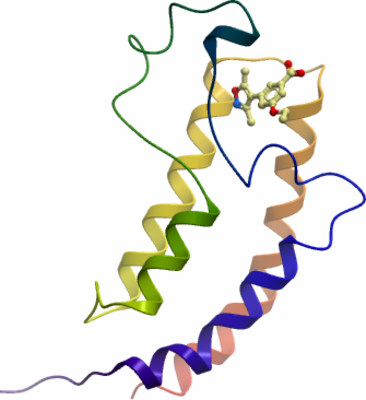
Crystallization: The protein (9.6 mg/ml) in SEC buffer was mixed with an equal volume (100 nl) of reservoir solution (0.2M Potassium thiocyanate; 20% (w/v) PEG3350; 5% (v/v) ethylene glycol and equilibrated as a sitting drop at 4 C in the presence of 10mM of E08681a (3-(3,5-dimethyl-isoxazol-4-yl)-5-ethoxy-benzoic acid)
Data Collection: Data were collected in-house on a Rigaku FRE SuperBright with an RAXIS-VI detector at a wavelength of 1.52 Å
Data Processing: Indexing and integration was carried out using MOSLFM and scaling was performed with SCALA. Initial phases were calculated by molecular replacement with PHASER3 using the previously determined model of CREBBP (PDB ID: 3DWY). Initial models were built by ARP/wARP followed by manual building in COOT. Refinement was carried out in REFMAC5 to 1.8 Å. Thermal motions were analyzed using TLSMD and hydrogen atoms were included in late refinement cycles.