Molecular Biology
Entry Clone Accession: IMAGE:4130242
Entry Clone Source: MGC
SGC Construct ID: FEVA-c012
Protein Region: K42-H141
Vector: pNIC28-Bsa4, N-terminal His tag followed by a TEV cleavage site.
Host: BL21(DE3)-R3-pRARE2
Sequence (with tag(s)): MHHHHHHSSGVDLGTENLYFQSMKGSGQIQLWQFLLELLADRANAGCIAWEGGHGEFKLTDPDEVARRWGERKSKPNMNYDKLSRALRYYYDKNIMSKVHGKRYAYRFDFQGLAQACQPPPAH
Sequence after tag cleavage: SMKGSGQIQLWQFLLELLADRANAGCIAWEGGHGEFKLTDPDEVARRWGERKSKPNMNYDKLSRALRYYYDKNIMSKVHGKRYAYRFDFQGLAQACQPPPAH
DNA Sequence: CATATGCACCATCATCATCATCATTCTTCTGGTGTAGATCTGGGTACCGAGAACCTGTACTTCCAATCCATGAAAGGCAGCGGACAGATCCAGCTGTGGCAGTTTCTGCTGGAGCTGCTGGCTGACCGCGCGAACGCCGGCTGCATCGCGTGGGAGGGCGGTCACGGCGAGTTCAAGCTCACGGACCCGGACGAGGTGGCGCGGCGGTGGGGCGAGCGCAAGAGCAAGCCCAACATGAACTACGACAAGCTGAGCCGCGCCCTGCGCTACTACTACGACAAGAACATCATGAGCAAGGTGCATGGCAAGCGCTACGCCTACCGCTTCGACTTCCAGGGCCTGGCGCAGGCCTGCCAGCCGCCGCCCGCGCACTGACAGTAAAGGTGGATACGGATCCGAA
Protein Expression
Medium: Terrific Broth
Antibiotics: Kanamycin
Procedure: Recombinant protein expression was induced by the addition of 0.1 mM isopropyl 1-thio-- D-galactopyranoside to bacterial cultures grown in TB (Terrific Broth) containing 50 mg/ml kanamycin at an OD600 of 3.0 at 37 °C in UltraYield baffled flasks (Thomson Instrument Co, Oceanside, CA). Cultures were further incubated at 18 °C overnight.
Protein Purification
Procedure: Cell pellets were thawed and resuspended in buffer A (50 mM HEPES, pH 7.5, 500 mM NaCl, 5% glycerol, 10 mM imidazole, 0.5 mM tris(2-carboxyethyl)phosphine (TCEP)), with the addition of 1x protease inhibitor set VII (Merck) and 15 units/ml Benzonase. Cells were lysed using sonication. Cell debris and nucleic acids were removed by addition of 0.15% polyethyleneimine, pH 7.5, and centrifugation at 40,000 g for 1 h at 4 °C. Clarified lysates were applied to a 3-ml Ni-iminodiacetic acid-immobilized metal ion affinity chromatography gravity flow column, washed with 20 column volumes (CV) of buffer A, followed by 20 CV of wash buffer (buffer A with 30 mM imidazole). Fractions were eluted in 2-CV aliquots of buffer A containing 300 mM imidazole and analyzed by SDS-PAGE, and relevant fractions pooled and cleaved with His6-tagged TEV protease (1:20 mass ratio) overnight at 8 °C. Imidazole was removed by concurrent dialysis during cleavage, using a 3.5-kDa MWCO snakeskin membrane in buffer B (20 mM HEPES, pH 7.5, 500 mM NaCl, 5% glycerol, 0.5 mM TCEP). TEV protease was removed from dialyzed proteins using NiIDA immobilized metal ion affinity chromatography (2-ml CV) and washed with an imidazole gradient in 20 mM steps to 100 mM in buffer B, and cleaved protein was pooled and concentrated with a 3-kDa MWCO centrifugal concentrators. Final separation was by size exclusion chromatography, using a HiLoad 16/60 Superdex S75 or 200 column equilibrated in buffer B, and run at 1.2 ml/min in buffer B.
Protein identity was confirmed by LC/ESI-TOF mass spectrometry with a single peak of mass 11794.5 corresponding to an expected mass of 11794.4.
Structure Determination
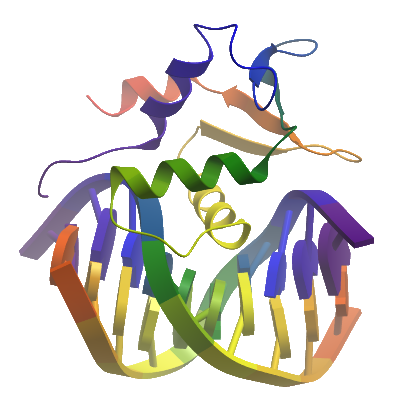
Crystallization: The oligonucleotides 5'-ACCGGAAGTG and 5'-CACTTCCGGT were resuspended to 1000uM in 10mM Tris-HCl -pH 8.0 and 250ul of each mixed with 50ul 10x annealing buffer (1x is 10mM Tris pH 7.5, 50mM NaCl) by heating to 95°C for 5 min & cooling to RT overnight in a heating block. Frozen protein (at 19mg/ml) mixed with dsDNA in a 1:1.1 ratio and incubated on ice for 20 minutes fro complex to bind. The DNA complex was concentrated approx. 6-fold in a 3kDa mwco concentrator. Final concentration of protein in dsDNA complex was approximately 16 mg/ml. Crystals were grown by vapor diffusion at 20°C in 150nl sitting drops. A crystal was grown by mixing 100nl of protein-dsDNA complex (16 mg/ml) and 50nl of mother liquor consisting of 25%(w/v) PEG 3350, 0.1M bis-tris pH 5.5. Crystals were loop mounted and transferred to a cyro solution consisting of well solution supplemented with the addition of 25 % v/v Ethylene glycol, before being flash cooled by being plunged directly into a pool of liquid nitrogen.
Data Collection: Diffraction data were collected on beamline I04 at Diamond Light Source, at a wavelength of 0.975Å to a maximum resolution of 2.0Å. The data was processing using the program XDS.
Data Processing: The structure was solved by Molecular replacement using the program PHASER and the coordinates of apo-FEV (2YPR) as a search model. DNA was modelled manually in COOT and the structure was refined using PHENIX REFINE to a final Rfactor= 21.2%, Rfree= 25.4%.