Entry Clone Source: MGC |
Entry Clone Accession: IMAGE:3686411 |
SGC Construct ID: pNIC-CTHF |
Vector: pNIC_CTHF. Details [PDF ]; Sequence [ FASTA ] or [ GenBank ] |
Final protein sequence (Tag sequence in lowercase):
MKEDLNLRLKKLTHAAPCMLFMKGTP
QEPRCGFSKQMVEILHKHNIQFSSFD
IFSDEEVRQGLKAYSSWPTYPQLYVS
GELIGGLDIIKELEASEELDTICPKA
aenlyfq^shhhhhhdykddddk
^ TEV cleavage site |
Tags and additions: TEV protease cleavable C-terminal histidine tag |
Host: BL21 (DE3)R3-pRARE2 (Phage resistant strain). |
Expression: TB supplemented with 50 µg/ml Kanamycin and 34 µg/ml chloramphenicol. 4 litre TB in two 4L flasks were innoculated with 40 ml overnight culture and grown at 37°C until OD600 reached 2.5. The temperature was then decreased to 18°C and the protein expression induced with 0.1 mM IPTG over night |
Cell harvest: The cells were collected by centrifugation (4000 RPM, 30 minutes) and frozen at -80°C |
Cell Lysis: Cell pellets were dissolved in approximately 150ml lysis buffer and broken by homogenization by 5 passes at 12,000 psi. The cell debris was pelleted at 35,000 x g and the supernatant used for further purification
Lysis buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 10 mM Imidazole, 5% glycerol, 0.5 mM TCEP, 1 mM PMSF and 3 U/ml of Benzonase
Extraction buffer, extraction method: The cell pellet was resuspended in a total volume of 200 ml lysis buffer and the cells disrupted by sonication. Nucleic acids and cell debris were removed by adding 0.15% PEI, followed by centrifugation for 60 minutes at 40,000xg. The supernatant was further clarified by filtration (0.20 mm). |
Column 1: Ni-sepharose, HisTrap FF, 5 ml (GE healthcare) |
Column 1 Buffers:
Binding buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 10 mM Imidazole, 0.5 mM TCEP
Wash and elution buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 10-300 mM imidazole |
Column 1 Procedure: The cell lysate was applied onto a 5 ml Ni-Sepharose FF gravity column equilibrated with binding buffer. The column was subsequently washed with 10 column volumes of binding buffer and the protein eluted using a stepwise gradient of imidazole. All fractions were collected and analysed by 4-12% SDS-PAGE. |
Column 2: Hiload 16/60 Superdex 200 prep grade 120 ml (GE/Amersham Biosciences). |
Column 2 Buffer:10 mM HEPES, pH 7.5, 500 mM NaCl, 5% glycerol, 0.5 mM TCEP |
Column 2 Procedure: The fractions containing TXNL2 from the Ni-affinity chromatography were concentrated in amicon (3K) to 4ml. The protein was filtered through a 0.20µm PVDF filter and applied onto a Superdex S200 column at 1.2 ml/min. The eluted proteins were collected in 1.8 ml fractions and analysed by SDS-PAGE.
|
Enzymatic treatment: TEV cleavage |
Column 3: Ni-Sepharose FF (TEV clean up) |
Column 3 Wash Buffer:10 mM HEPES, pH 7.5, 500 mM NaCl, 5% glycerol |
Column 3 Procedure: The N-terminal histidine-tag was cleaved with 150 µg of TEV protease per 10 mg protein at 4°C for 48 hours. The protein was then applied to a 0.5 ml Ni-sepharose column and the flow through collected. The column washed with 2.5 ml buffer and the flow through and the wash fraction analysed by SDS-PAGE.
|
Concentration: The protein was concentrated in Amicon 3K to 36.8 mg/ml. The protein concentration was determined spectrophotometrically using the predicted molar extinction coefficient 11460(M-1 cm-1) and the predicted mass of 12727.7 Da |
Mass spectrometry characterization:
Observed mass: 12728.1 Da
Expected mass: 12727.7 Da
|
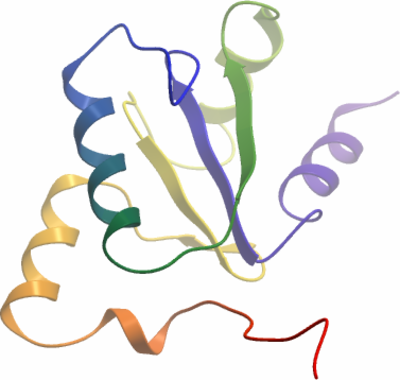
Crystallisation: TXNL2A was crystallised by vapor diffusion at 4°C from a sitting drop consisting of 50 nl protein (36.8 mg/ml) and 100 nl well solution containing 0.1 M Bis-Tris pH 5.5, 25% PEG3350. The crystal was transferred to a cryo protectant composed of 25% ethylene glycol before flash-cooling in liquid nitrogen |
Data collection:
Resolution: 1.8Å.
X-ray source: Diamond Light Source beamline I03. |