Vector: pNIC-Zb.
|
Amplified DNA sequence:
TACTTCCAATCCATGCCGAAGCTGCC
TGAGAATTATACCGATGAAACCTGGC
AGAAACTGAAGGAAGCCGTGGAAGCA
ATCCAGAACAGTACCTCGATTAAATA
CAATCTGGAAGAGCTGTATCAGGCCG
TCGAGAACCTGTGCTCCTACAAAATC
TCTGCTAATCTGTATAAGCAGCTGCG
TCAGATTTGTGAGGACCACATCAAGG
CGCAGATTCATCAGTTCCGCGAAGAC
AGCCTGGATTCCGTGCTGTTTCTGAA
GAAAATCGATCGTTGCTGGCAGAACC
ACTGTCGTCAGATGATCATGATTCGC
TCAATTTTCCTGTTTCTGGACCGCAC
CTACGTCCTGCAGAACAGCATGCTGC
CGAGTATCTGGGATATGGGTCTGGAA
CTGTTCCGTGCCCATATCATTTCTGA
CCAGAAAGTTCAGAACAAGACCATCG
ATGGTATTCTGCTGCTGATCGAGCGT
GAACGCAATGGCGAGGCTATTGACCG
TTCCCTGCTGCGCTCTCTGCTGTCAA
TGCTGAGTGACCTGCAGATTTACCAG
GATTCTTTCGAACAGCGTTTTCTGGA
AGAGACCAACCGCCTGTATGCCGCTG
AGGGCCAGAAACTGATGCAGGAGCGC
GAAGTTCCTGAATACCTGCACCATGT
GAATAAGCGTCTGGAAGAGGAAGCCG
ACCGCCTGATCACCTACCTGGATCAG
ACCACCCAGAAAAGCCTGATTGCGAC
CGTGGAGAAGCAGCTGCTGGGTGAAC
ACCTGACCGCAATCCTGCAGAAAGGC
CTGAACAATCTGCTGGACGAGAACCG
TATTCAGGATCTGTCGCTGCTGTATC
AGCTGTTCAGCCGTGTCCGCGGTGGC
GTGCAGGTCCTGCTGCAGCAGTGGAT
CGAATACATTAAGGCGTTTGGCAGCA
CCATCGTTATTAACCCGGAGAAAGAC
AAGACCATGCGCCAGGAACTGGACGA
TTTCAAAGACAAGGTGGATCATATCA
TTGATATCTGCTTCCTGAAGAACGAA
AAGTTCATCAACGCCATGAAGGAAGC
CTTCGAAACCTTTATCAACAAACGTC
CCAATTGACAGTAAAGGTGGATA
|
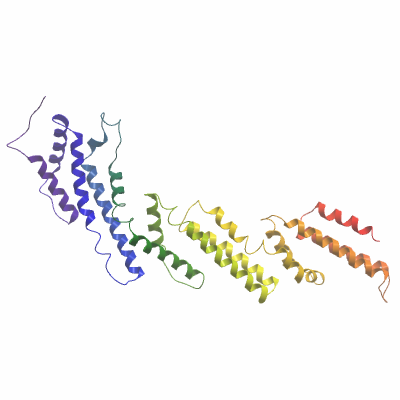
Final protein sequence (Tag sequence in lowercase):
mhhhhhhssgvdnkfnkerrrarrei
rhlpnlnreqrrafirslrddpsqsa
nllaeakklndaqpkgtenlyfq^sM
PKLPENYTDETWQKLKEAVEAIQNST
SIKYNLEELYQAVENLCSYKISANLY
KQLRQICEDHIKAQIHQFREDSLDSV
LFLKKIDRCWQNHCRQMIMIRSIFLF
LDRTYVLQNSMLPSIWDMGLELFRAH
IISDQKVQNKTIDGILLLIERERNGE
AIDRSLLRSLLSMLSDLQIYQDSFEQ
RFLEETNRLYAAEGQKLMQEREVPEY
LHHVNKRLEEEADRLITYLDQTTQKS
LIATVEKQLLGEHLTAILQKGLNNLL
DENRIQDLSLLYQLFSRVRGGVQVLL
QQWIEYIKAFGSTIVINPEKDKTMRQ
ELDDFKDKVDHIIDICFLKNEKFINA
MKEAFETFINKRPN
^ TEV cleavage site |
Tags and additions: Tev-cleavable (^) N-terminal hexahistidine and Z tag. |
Host: BL21 (DE3)R3-pRARE2 Phage-resistant derivative of BL21 (DE3), with pRARE2 plasmid encoding rare codon tRNAs (chloramphenicol-resistant).
|
Growth medium, induction protocol: A glycerol stock was used to inoculate a 10ml starter culture containing 2YT media with 50µg/ml Kanamycin and 34µg/ml Chloramphenicol. The starter culture was grown overnight at 37°C with shaking at 200 rpm. The following morning, four flasks containing 1 L LB/Kanamycin were each inoculated with 1 ml of the starter culture. Cultures were incubated at 37°C with shaking at 180 rpm until an OD600nm = 0.4 was reached. The flasks were then cooled down to 18°C to an OD600nm = 0.7 and added 0.4mM IPTG to induce protein expression overnight. Cells were harvested by centrifugation at 4500 rpm at 4°C for 15 min. Cell pellets from each flask were resuspended in 25ml binding buffer (50mM HEPES, pH 7.5; 500mM NaCl; 5% Glycerol; 5mM Imidazole).
Extraction buffer, extraction method: The cells were lysed by ultrasonication over 7 min with the sonicator pulsing ON for 5 sec and OFF for 10 sec. The cell lysate was spun down by centrifugation at 18000 rpm at 4°C for 1 h. The supernatant was recovered for purification. |
Column 1: Ni-Affinity Chromatography twinned with DE-52 column 2.5g of DE-52 resin dissolved in 25ml 2.5M NaCl was applied onto a drip column and equilibrated with Binding buffer. 5ml of 50% Ni-IDA slurry was applied onto a 1.5 x 10cm column. The column was first washed with deionised distilled H2O, and then equilibrated with Binding buffer. DE-52 column was mounted on top of the Ni column. |
Column 1 Buffers:
Binding buffer: 50 mM HEPES, pH 7.5; 500 mM NaCl; 5% glycerol; 5 mM Imidazole.
Wash buffer: 50 mM HEPES, pH 7.5; 500 mM NaCl; 5% glycerol; 30 mM Imidazole.
Elution buffer: 50 mM HEPES, pH 7.5; 500 mM NaCl; 5% glycerol, 50 to 250 mM Imidazole. |
Column 1 Procedure: The supernatant was applied by gravity flow onto the DE-52 column and then passed onto the Ni column. The Ni column was washed with Wash buffer and the bound protein was eluted by applying a step gradient of Imidazole (5 ml fractions of Elution buffer supplemented with 50mM, 100mM, 150mM and 2x10ml fractions with 250mM Imidazole). 10mM L-Arginine and 10 mM L-Glutamic Acid were added to each fraction collected for overnight storage at 4°C. |
Enzymatic treatment:TEV protease cleavage. Fractions containing CUL4B were treated with TEV protease overnight at 4°C. |
Column 2: Reverse Purification Ni-Affinity Chromatography 500µl of 50% Ni-IDA slurry was applied onto a BioRad Poly-Prep disposable drip column. The column was first washed with deionised distilled H2O, and then equilibrated with Binding buffer. |
Column 2 Buffers:
Elution buffer: 50 mM HEPES, pH 7.5; 500 mM NaCl; 5% glycerol, 250 mM Imidazole.
|
Column 2 Procedure: The CUL4B pool treated with TEV was applied by gravity flow onto the Ni-IDA column. The flow through was collected, washed the column with 10ml Wash buffer and the bound impurities were eluted with 500µl of elution buffer with 250mM Imidazole. |
Column 3: Anion Exchange Chromatography MonoQ 5/50 GL (GE Healthcare) |
Column 3 Buffers:
IEX Buffer I: 50 mM HEPES, pH 7.5;
IEX Buffer II: 1M NaCl, 50mM HEPES pH 7.5.
|
Column 3 Procedure: The MonoQ 5/50 column was first washed with IEX buffer II and then equilibrated with IEX buffer I. The CUL4B protein in the flow through of column 2 was concentrated to 5ml using an Amicon Ultra-15 filter with a 30kDa cut-off. The 5ml fraction was made up to 50ml with IEX buffer I and applied onto the column. Bound protein was eluted in 0%-100% gradient with IEX buffer II. Clean fractions containing the protein were pooled together. |
Column 4: Reverse Purification Ni-Affinity Chromatography 500µl of 50% Ni-IDA slurry was applied onto a BioRad Poly-Prep disposable drip column. The column was first washed with deionised distilled H2O, and then equilibrated with Binding buffer. |
Column 4 Buffers:
Binding buffer: 50mM HEPES, pH 7.5; 500mM NaCl; 5% Glycerol; 5mM Imidazole.
Wash buffer: 50mM HEPES, pH 7.5; 500mM NaCl; 5% Glycerol; 30mM Imidazole.
Elution buffer: 50mM HEPES, pH 7.5; 500mM NaCl; 5% Glycerol; 250mM Imidazole.
|
Column 4 Procedure: Protein from Anion exchange step was applied by gravity flow onto the Ni-IDA column. The flow through was collected; after a wash step with 10ml Wash, buffer bound impurities were eluted with 500µl of elution buffer with 250mM Imidazole. 10mM DTT and 10mM L-Arginine/L-Glutamic Acid were added to the flow through containing CUL4B. |
Protein concentration: The protein was concentrated in an Amicon Ultra-4 filter with a 30 kDa cut-off. |
Mass spectrometry characterization: The purified native protein was homogeneous and had an experimental mass after tag cleavage of 41901.1, just above the expected mass of 41898.3Da. Masses were determined by LC-MS, using an Agilent LC/MSD TOF system with reversed-phase HPLC coupled to electrospray ionization and an orthogonal time-of-flight mass analyzer. |
Crystallisation: Protein was buffered in 175mM NaCl, 50mM HEPES pH 7.5, 10mM DTT, 10mM L-Arginine/L-Glutamic Acid. Protein was concentrated to 11mg/ml (calculated using extinction co-efficient of 39880). Native crystals were grown at 20°C in 150nl sitting drops mixing 75nl protein solution with 75nl of a reservoir solution containing 24% PEG 4000, 0.1M Li2SO4, 100mM Tris pH 8.2. On mounting crystals were cryo-protected with an additional 25% Ethylene glycol. |
Data collection:
X-ray source: Diamond I02
Resolution: 2.6Å |