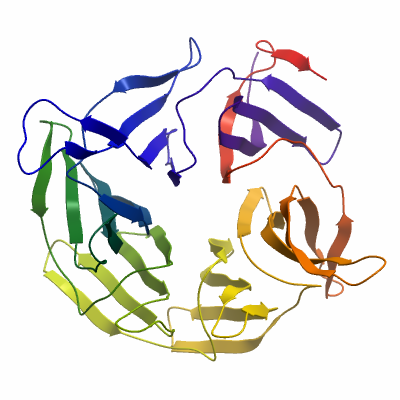
KBTBD5A (4ASC) Materials & Methods |
Entry Clone Source: MGC |
SGC Construct ID: KBTBD5A-c006 |
Vector: pNIC-CTHF. Details [PDF ]; Sequence [ FASTA ] or [ GenBank ]. T7/lac regulated, C-terminal His-tag and Flag tag, TEV, LIC cloning using BfuAI /T4 polymerase, pET28 backbone. |
DNA sequence:
TTAAGAAGGAGATATACTATGTTCCTGCA
GGATCTCATCTTCATGATCAGTGAGGAGG
GCGCTGTGGCCTACGATCCAGCAGCCAAC
GAGTGCTACTGTGCTTCCCTCTCCAGCCA
GGTCCCCAAGAACCACGTCAGCCTGGTTA
CCAAGGAGAACCAGGTCTTCGTGGCTGGA
GGCCTCTTCTACAACGAAGACAACAAAGA
GGACCCCATGAGCGCATACTTCCTGCAGT
TTGACCATCTGGACTCAGAGTGGCTGGGG
ATGCCACCGCTGCCCTCGCCCCGCTGCCT
CTTTGGCCTGGGAGAAGCTCTCAACTCCA
TCTACGTGGTCGGTGGCAGAGAGATCAAG
GACGGCGAGCGCTGCCTGGACTCGGTCAT
GTGCTACGACAGGCTGTCATTCAAATGGG
GTGAATCGGACCCGCTGCCTTACGTGGTG
TATGGCCACACAGTGCTCTCCCACATGGA
CCTTGTCTACGTAATTGGCGGCAAAGGCA
GTGACAGGAAGTGCCTGAACAAGATGTGC
GTCTATGACCCCAAGAAGTTTGAGTGGAA
GGAGCTGGCACCCATGCAGACCGCCCGCT
CACTCTTTGGGGCCACTGTCCATGATGGC
CGCATTATCGTGGCAGCTGGGGTCACCGA
CACAGGGCTGACCAGTTCTGCCGAAGTGT
ACAGCATCACAGACAACAAGTGGGCACCC
TTCGAGGCCTTCCCACAGGAGCGTAGCTC
ACTCAGCCTGGTCAGCCTGGTGGGTACCC
TCTATGCCATTGGTGGCTTTGCCACACTG
GAGACGGAGTCTGGAGAGCTGGTTCCCAC
AGAGCTCAATGACATCTGGAGGTATAACG
AGGAGGAGAAGAAATGGGAGGGTGTCCTG
CGGGAGATCGCCTATGCAGCAGGTGCCAC
CTTCCTACCAGTGCGGCTCAATGTGCTGC
GCCTGACTAAGATGGCAGAGAACCTCTAC
TTCCAATC
|
Final protein sequence (Tag sequence in lowercase):
MFLQDLIFMISEEGAVAYDPAANECY
CASLSSQVPKNHVSLVTKENQVFVAG
GLFYNEDNKEDPMSAYFLQFDHLDSE
WLGMPPLPSPRCLFGLGEALNSIYVV
GGREIKDGERCLDSVMCYDRLSFKWG
ESDPLPYVVYGHTVLSHMDLVYVIGG
KGSDRKCLNKMCVYDPKKFEWKELAP
MQTARSLFGATVHDGRIIVAAGVTDT
GLTSSAEVYSITDNKWAPFEAFPQER
SSLSLVSLVGTLYAIGGFATLETESG
ELVPTELNDIWRYNEEEKKWEGVLRE
IAYAAGATFLPVRLNVLRLTKMAenl
yfq^shhhhhhdykddddk
^ TEV protease recognition site
|
Tags and additions: Cleavable C-terminal His6 and FLAG tag |
Host: BL21(DE3)-R3-pRARE2. Phage-resistant derivative of BL21 (DE3), with pRARE2 plasmid encoding rare codon tRNAs (chloramphenicol-resistant). |
Growth Medium & Induction Protocol: A glycerol stock was used to inoculate a 10ml starter culture containing LB media with 50µg/ml Kanamycin and 34µg/ml Chloramphenicol. The starter culture was grown overnight at 37°C with shaking at 200 rpm. The following morning, six flasks containing 1L LB/Kanamycin were each inoculated with 1 ml of the starter culture. Cultures were incubated at 37°C with shaking at 180 rpm until an OD600nm = 0.4 was reached. The flasks were then cooled down to 18°C to an OD600nm = 0.7 and 0.5mM IPTG added to induce protein expression overnight. Cells were harvested by centrifugation in a JLA 8.1 rotor at 4000 rpm at 4°C for 15 min. Cell pellets from each tube were resuspended in 25ml Binding buffer (50mM HEPES, pH 7.5; 500mM NaCl; 5% Glycerol; 5mM Imidazole) with 0.5mM TCEP, 1mM PMSF and 0.1% DDM.
|
Extraction buffer, extraction method: The cells were lysed by ultrasonication over 10min with the sonicator pulsing ON for 5 sec and OFF for 10min. The cell lysate was spun down by centrifugation in JA 18 rotor at 18000 rpm at 4°C for 1 h. The supernatant was recovered for purification.
|
Column 1: Ni-Affinity Chromatography twinned with DE-52 column
2.5g of DE-52 resin dissolved in 25ml 2.5M NaCl was applied onto a drip column and equilibrated with Binding buffer. 5ml of 50 % Ni-IDA slurry was applied onto a 1.5 x 10 cm column. The column was first washed with deionised distilled H2O, and then equilibrated with Binding buffer. DE-52 column was mounted on top of the Ni column. |
Column 1 Buffers:
Binding buffer: 50mM HEPES, pH 7.5; 500mM NaCl; 5% Glycerol; 5mM Imidazole
Wash buffer: 50mM HEPES, pH 7.5; 500mM NaCl; 5% Glycerol; 30mM Imidazole
Elution buffer: 50mM HEPES, pH 7.5; 500mM NaCl; 5% Glycerol; 50 to 250mM Imidazole
|
Column 1 Procedure: The supernatant was applied by gravity flow onto the DE-52 column to remove nucleic acids and then passed onto the Ni column. The Ni column was washed with Wash buffer and the bound protein was eluted by applying a step gradient of Imidazole (5 ml fractions of Elution buffer supplemented with 50mM, 100mM, 150mM and 2x10ml fractions with 250mM Imidazole). 10mM Arginine/10 mM Glutamate mix and 10mM DTT was added to each fraction collected for overnight storage at 4°C.
|
Enzymatic Treatment: TEV protease cleavage. Fractions containing KBTBD5 were treated with TEV protease overnight at 4°C.
|
Column 2: Size Exclusion Chromatography HiLoad 16/60 Superdex S75 (GE Healthcare). |
Column 2 Buffers:
Gel Filtration buffer: 300mM NaCl, 50mM HEPES pH 7.5, 0.5mM TCEP
|
Column 2 Procedure: The Superdex S75 column was first equilibrated with Gel Filtration buffer. Concentrated the protein fraction from above step to
|
Column 2: Anion Exchange Chromatography HiTrap Q HP 5ml column |
Column 2 Buffers:
IEX buffer A (binding): 50mM HEPES pH 7.5
IEX buffer B (elution): 1M NaCl, 50mM HEPES pH 7.5
|
Column 2 Procedure: The HiTrap Q HP column was first washed with IEX buffer II and then equilibrated with IEX buffer I. The protein fraction from above step was made up to 90ml with IEX buffer I and applied onto the column connected to an Äkta Express. Bound protein was eluted in 0%-100% gradient with IEX buffer II. Fractions containing the pure protein were pooled together.
|
Concentration: The protein was concentrated in an Amicon Ultra-4 30kDa MWCO filter. |
Mass spec characterization: The purified native protein produced experimental masses after tag cleavage of 35205 Da and 35222Da. The smaller mass matches precisely the expected MW, while the 17Da higher peak which was more populated appears to represent oxidation. Masses were determined by LC-MS, using an Agilent LC/MSD TOF system with reversed-phase HPLC coupled to electrospray ionization and an orthogonal time-of-flight mass analyzer. |
Crystallization: Protein was buffered in 50mM HEPES pH 7.5, 120mM NaCl, 10mM DTT, 10mM Arginine, 10mM Glutamate. Protein was concentrated to 11mg/ml (calculated using extinction co-efficient of 55350). Native crystals were grown at 4°C in 150nl sitting drops mixing 100nl protein solution with 50nl of a reservoir solution containing 20% (w/v) PEG 6000, 0.1M Citrate pH 5.3. On mounting crystals were cryo-protected with an additional 25% Ethylene glycol. |
Data Collection:
Resolution: 1.8Å X-ray source: Diamond IO4-1
Crystals of KBTBD5A diffracted to a resolution of 1.8 Å (scaled resolution). A full dataset was collected at 100 K on Diamond Light Source beamline I04-1. Crystals belonged to the Orthorhombic space group P212121 with unit-cell parameters a=61.33 Å b=64.95 Å c=89.2 Å, α=90° β= 90° γ= 90°. Only one molecule was present in the asymmetric unit. Data were indexed and integrated using iMOSFLM and scaled using AIMLESS. Phases were found using molecular replacement in PHASER. PHENIX.SCULPTOR was used to optimize the PDB entry 2WOZ for use as a search model. An initial model was built using PHENIX.AUTOBUILD and the structure refined and modified using alternate rounds of REFMAC5 and COOT. The final model was validated using the JCSG Quality Control Server.
|