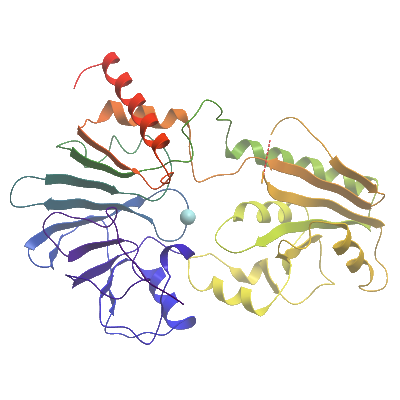
Molecular Biology
Entry Clone Accession: IMAGE:3048375
Entry Clone Source: MGC
SGC Construct ID: DCLRE1AA-c290
Protein Region: A676-Y1040
Vector: pFB-LIC-Bse. This is a baculovirus transfer vector (Bac-to-bac), with N-terminal 6 His tag followed by a TEV cleavage site
Host: DH10Bac
Sequence (with tag(s)): MGHHHHHHSSGVDLGTENLYFQSMAHGGLQRGNKKIPESSNVGGSRKKTCPFYKKIPGTGFTVDAFQYGVVEGCTAYFLTHFHSDHYAGLSKHFTFPVYCSEITGNLLKNKLHVQEQYIHPLPLDTECIVNGVKVVLLDANHCPGAVMILFYLPNGTVILHTGDFRADPSMERSLLADQKVHMLYLDTTYCSPEYTFPSQQEVIRFAINTAFEAVTLNPHALVVCGTYSIGKEKVFLAIADVLGSKVGMSQEKYKTLQCLNIPEINSLITTDMCSSLVHLLPMMQINFKGLQSHLKKCGGKYNQILAFRPTGWTHSNKFTRIADVIPQTKGNISIYGIPYSEHSSYLEMKRFVQWLKPQKIIPTVNVGTWKSRSTMEKYFREWKLEAGY
Sequence after tag cleavage: SMAHGGLQRGNKKIPESSNVGGSRKKTCPFYKKIPGTGFTVDAFQYGVVEGCTAYFLTHFHSDHYAGLSKHFTFPVYCSEITGNLLKNKLHVQEQYIHPLPLDTECIVNGVKVVLLDANHCPGAVMILFYLPNGTVILHTGDFRADPSMERSLLADQKVHMLYLDTTYCSPEYTFPSQQEVIRFAINTAFEAVTLNPHALVVCGTYSIGKEKVFLAIADVLGSKVGMSQEKYKTLQCLNIPEINSLITTDMCSSLVHLLPMMQINFKGLQSHLKKCGGKYNQILAFRPTGWTHSNKFTRIADVIPQTKGNISIYGIPYSEHSSYLEMKRFVQWLKPQKIIPTVNVGTWKSRSTMEKYFREWKLEAGY
DNA Sequence: CCATGGGCCACCATCATCATCATCATTCTTCTGGTGTAGATCTGGGTACCGAGAACCTGTACTTCCAATCCATGGCTCATGGTGGGCTGCAAAGGGGCAACAAGAAAATCCCAGAGTCATCTAATGTAGGAGGATCAAGAAAAAAGACATGTCCATTCTATAAGAAAATACCTGGAACCGGCTTTACAGTTGATGCCTTTCAGTATGGCGTGGTTGAAGGTTGCACAGCCTATTTTCTCACACATTTTCATTCTGATCATTATGCTGGATTGTCTAAACACTTCACATTTCCAGTTTATTGTAGTGAGATAACTGGCAATTTGTTGAAGAACAAGCTTCATGTGCAAGAACAATATATTCACCCATTGCCACTGGACACTGAATGTATTGTGAATGGTGTCAAAGTTGTTTTGCTTGATGCCAATCACTGTCCAGGTGCTGTCATGATCCTCTTTTATCTTCCTAATGGTACTGTCATATTACACACGGGAGACTTCAGAGCAGATCCCAGCATGGAACGTTCTCTTCTTGCGGACCAGAAAGTCCATATGCTGTACTTAGATACCACATATTGTAGCCCAGAATACACCTTTCCATCTCAGCAAGAGGTTATCCGGTTTGCCATCAACACTGCCTTTGAGGCTGTAACTCTAAACCCACATGCTCTTGTTGTCTGTGGCACTTACTCTATTGGAAAAGAGAAAGTCTTCCTAGCCATTGCTGATGTTTTAGGTTCAAAAGTGGGCATGTCCCAGGAAAAATATAAAACTCTACAGTGCCTCAATATACCAGAAATTAATTCACTCATCACTACCGACATGTGCAGTTCATTGGTTCACCTTCTCCCAATGATGCAAATTAATTTTAAGGGCTTACAGAGTCATTTGAAGAAGTGTGGTGGGAAATACAATCAGATTTTGGCATTTCGACCTACAGGATGGACACACTCTAACAAGTTCACTAGAATAGCAGATGTTATTCCCCAGACCAAAGGAAACATTTCAATATATGGAATTCCTTACAGTGAACACAGCAGCTACCTAGAAATGAAGCGCTTTGTCCAGTGGCTGAAGCCCCAGAAAATCATACCTACTGTAAATGTGGGCACCTGGAAATCTAGGAGCACAATGGAGAAATATTTTAGAGAGTGGAAATTGGAAGCTGGATATTGACAGTAAAGGTGGATACGGATCCGAATTCGAGCTCCGTCGACAAGCTT
Protein Expression
Medium: SF900II
Antibiotics: Ampicillin
Procedure: Baculoviruses were generated by recombination in E. coli DH10Bac (Life Technologies) followed by transfection and two rounds of amplification in SF9 cells. DCLRE1A was expressed in 1-L cultures of SF9 cells in 4-L shaker flasks at 27°C, infected at 2 × 106 cells/ml with 3 ml of virus, and incubated for further 70 h. The cells were collected by centrifugation, suspended in 15 ml/l of lysis buffer (50 mM HEPES, pH 7.5, 0.5 M NaCl, 5% v/v glycerol, 10 mM imidazole, and 1 mM TCEP) and frozen at −80°C.
Protein Purification
Procedure: Cells were thawed, 3–5 volumes of lysis buffer were added, and the cells were disrupted by sonication. The lysate was centrifuged for 30 min at 40 000 × g, and the clear supernatant was collected. The clarified cell lysate was loaded on a 5-ml Histrap FF column. The column was washed with 20 volumes of wash buffer (lysis buffer with 30 mM imidazole), and the protein was recovered with elution buffer (lysis buffer with 300 mM imidazole). The eluted protein was combined with His10-tagged TEV protease (1/10 w/w) in a dialysis tubing, and digestion of the tag was performed overnight at 6°C while dialysing against 4 l of wash buffer. The material was then passed through a 1-ml HisTrap column to remove contaminating proteins and remaining TEV protease. The column was developed with a 40-ml gradient from wash buffer to elution buffer, and all fractions were analyzed by SDS-PAGE.
The DCLRE1A containing fractions from the second IMAC column were combined, concentrated to
The protein was analyzed by ESI-TOF intact mass spectrometry, with a significant peak at mass of 39117.9 Da being observed in addition to the expected mass of 41324 Da. Later analysis revealed that the crystallized molecule corresponded to the mass of 39117.9 Da which corresponds to a fragment spanning residues 696-1040, this interpretation was conserved by mass spectrometry of washed crystals.
Structure Determination
Crystallization: Protein crystallization was performed by vapour diffusion in sitting drops at 4°. A protein solution at 10 mg/mL was mixed at with a 2:1 ratio of protein to crystallization solution containing 23% PEG 3350, 0.1M bis-tris-propane pH 7.5, 0.2M sodium iodide, 5% ethylene glycol. Crystals were transferred to a cryo protectant solution consisting of well solution supplemented with an additional 20 % Ethylene Glycol before being loop mounted and plunged directly into a pool of liquid nitrogen. For structure solution a Platinum derivative crystal was prepared by soaking a crystal in a solution of 2.5 mM Potassium Tetrachloroplatinate for approximately 1hr before harvesting and cryo cooling as above.
Data Collection: Derivative data was collected at Diamond Light Source beamline I02 to 2.6Å resolution and a native data set was collected at Diamond Light Source beamline I02 to 2.16Å resolution and processed using MOSFLM.
Data Processing: The structure was solved by single isomorphous replacement with anomalous scattering (SIRAS) using the programs SHARP and SHELX. Two platinum ions were found and the density modified map produced at 2.5 Å allowed tracing of the entire molecule. The initial model was then used to solve the 2.16 Å dataset by molecular replacement using MOLREP. Refinement was performed using REFMAC to a final Rfactor = 17.6%, Rfree = 21.8%.