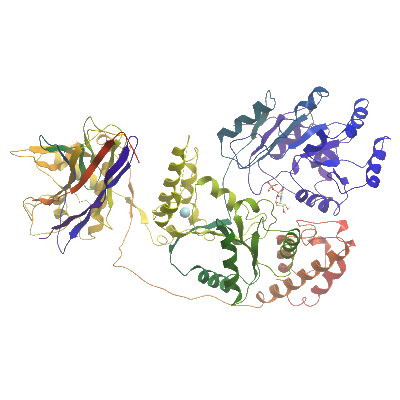
BLM
PDB:4CDG
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:Entry Clone Source:SGC Clone Accession:BLMA-c800, YYBLMA-c075
Tag:Tag TEV-cleavable N-terminal his6 tag, C-terminal non cleavable his6 tag.
Host:BL21(DE3)-R3-pRARE2
Construct
Prelude:Sequence:BLMmhhhhhhssgvdlgtenlyfq^SMSRNLKHERFQSLSFPHTKEMMKIFHKKFGLHNFRTNQLEAINAALLGEDCFILMPTGGGKSLCYQLPACVSPGVTVVISPLRSLIVDQVQKLTSLDIPATYLTGDKTDSEATNIYLQLSKKDPIIKLLYVTPEKICASNRLISTLENLYERKLLARFVIDEAHCVSQWGHDFRQDYKRMNMLRQKFPSVPVMALTATANPRVQKDILTQLKILRPQVFSMSFNRHNLKYYVLPKKPKKVAFDCLEWIRKHHPYDSGIIYCLSRRECDTMADTLQRDGLAALAYHAGLSDSARDEVQQKWINQDGCQVICATIAFGMGIDKPDVRFVIHASLPKSVEGYYQESGRAGRDGEISHCLLFYTYHDVTRLKRLIMMEKDGNHHTRETHFNNLYSMVHYCENITECRRIQLLAYFGENGFNPDFCKKHPDVSCDNCCKTKDYKTRDVTDDVKSIVRFVQEHSSSQGMRNIKHVGPSGRFTMNMLVDIFLGSKSAKIQSGIFGKGSAYSRHNAERLFKKLILDKILDEDLYINANDQAIAYVMLGNKAQTVLNGNLKVDFMETENSSSVKKQKALVAKVSQREEMVKKCLGELTEVCKSLGKVFGVHYFNIFNTVTLKKLAESLSSDPEVLLQIDGVTEDKLEKYGAEVISVLQKYSEWTSPAEDSSPGMSENLYFQSGGGLNDIFEAQKIEWHE
NanobodyMKYLLPTAAAGLLLLAAQPAMAQVQLQESGGGLVQAGGSLRLSCAASGIWFSINNMAWYRQTPGKQRERIAIITSAGTTNYVDSVKGRFTISRDDAKNTMYLQMNSLIPEDTAVYYCNLVADYDMGFQSFWGRGTQVTVSSHHHHHH
Vector:pNIC-NHT-CTB, pMES4 (amp) Details [
PDF ]; Sequence [
FASTA ] or [
GenBank ].
Growth
Medium:The expression plasmid was transformed into the host strain and plated on LB-agar containing 50 µg/ml kanamycin and 34 µg/ml chloramphenicol. Several colonies were combined to inoculate a 1ml culture in TB (+ 50 µg/ml kanamycin, 34 µg/ml chloramphenicol). The culture was grown overnight, glycerol was added to 15% v/v (from a 50% stock), and the resulting glycerol stock was frozen at -80°C in 100 µl aliquots. A loopful of cells from the glycerol stock was inoculated into 10-ml of TB medium containing 50 µg/ml kanamycin and 34 µg/ml chloramphenicol and grown overnight at 37°C. 1L of TB medium (containing 50 µg/ml kanamycin) was inoculated with 10 ml of the overnight culture and grown in 2 L UltraYield baffled flasks until OD600 of 2.0. Cells were cooled to 18°C, IPTG added to 0.1 mM and growth continued at 18°C overnight. The cells were collected by centrifugation then the pellets were scraped out and transferred to 50-ml Falcon tubes and frozen at -80°C.
Antibiotics:Procedure:Purification
BuffersProcedurePurification of Nanobody: Cell paste from each litre over expression was resuspended in 150ml of sucrose buffer (200mM Hepes pH7.5, 500mM Sucrose, 0.5mM EDTA) and left one ice for 1 hour with occasional stirring. 150ml of water was then added to each sample and left on ice for another hour with occasional stirring. The samples were then centrifuged for 40min at 5000 rpm and 10°C. 3.3ml of Ni-IDA beads were added to the supernatant and incubated for 1.5 hr. The resin was washed initially with 10 column volumes of lysis buffer and pelleted by centrifugation. Ten column volumes of wash buffer1 were added and the resuspended resin was transferred to a gravity flow column and washed with 5 column volumes of wash buffer. Proteins were eluted by addition of 2 x 10 ml of elution buffer.
Preparation of the BLM Nanobody complex: To form the BLM nanobody complex, BLM and Nanobody were mixed in a molar ratio of 1:1.1 and incubated on ice for 30 mins. The complex was loaded onto gel filtration Hiload 16/60 Superdex S200 prep grade, 120 ml in GF buffer, in order to separate the complex from unbound material.
Extraction
BuffersProcedureCell extraction : Frozen cell pellets were thawed briefly in a bath of warm water (20 - 37°C) then transferred to ice. Six volumes (i.e. 6 ml for every gram of cells) of lysis buffer was added, and the cells were resuspended by agitating and disrupted by pulsed sonication (5 sec on 10 sec off) for 15 minutes on ice. Nucleaic acids were removed by the addition of 0.1% Polyethylenimine and debris were removed by centrifugation for 25 minutes at 65,000xg.
Lysis buffer: 10 mM HEPES, pH 7.5, 500 mM NaCl, 5% glycerol, 10 mM Imidazole, 0.5 mM TCEP.
Column 1: 2ml bed volume Ni-IDA
Solutions:
Wash buffer: 50 mM HEPES, pH 7.5, 500 mM NaCl, 30 mM imidazole, 5% Glycerol, 0.5 mM TCEP.
Elution buffer: 50 mM HEPES, pH 7.5, 500 mM NaCl, 300 mM imidazole, 5% Glycerol, 0.5 mM TCEP.
Procedure: The cell extract was incubated with 5ml of Ni-IDA resin (GE Healthcare) for 1hr at 4°c with gentle agitation. The resin was pelleted by centrifugation (700g for 6 min), and the supernatant was decanted. The resin was washed initially with 10 column volumes of lysis buffer and pelleted by centrifugation. Ten column volumes of wash buffer1 were added and the resuspended resin was transferred to a gravity flow column and washed with 5 column volumes of wash buffer. Proteins were eluted by addition of 2 x 10 ml of elution buffer and combined with the fractions from the wash buffer.
Enzymatic treatment and Column 2: The N-terminal His6-tag was cleaved by incubating the protein overnight with tobacco etch virus (TEV) protease (1:40 ratio at 4°C) whilst being dialyzed into dialysis buffer (50 mM HEPES, 500 mM NaCl, 5 % Glycerol, 3 mM TCEP) using a 3,500 MWCO snakeskin dialysis membrane.
Cleaved protein was purified by passing over a 5 ml pre-equilibriated 50% Ni-IDA bead solution. Elution was done in GF buffer supplemented with step gradients of 20 mM, 40 mM 100 mM and 300 mM imidazole, with flowthrough and fractions collected.
Column 3 : Gel filtration, Hiload 16/60 Superdex S200 prep grade, 120 ml (GE Healthcare)
GF buffer: 10 mM HEPES, pH 7.5, 500 mM NaCl, 5 % Glycerol, 0.5 mM TCEP.
Procedure: The fractions from Ni-IDA column were concentrated to 2ml using a 50 KDa MWCO centrifugal concentrator and loaded on the gel filtration column in GF buffer at 1 ml/min. Eluted proteins were collected in 2-ml fractions and analysed on SDS-PAGE.
Concentration:The cleaved purified protein was concentrated using a 50 KDa MWCO centrifugal concentrator to 15 mg/ml and stored at 4°C. The protein concentration was determined spectrophotometrically using ε280 = 67730
LigandMassSpec:Crystallization:Crystallisation experiments were performed using the sitting drop vapour diffusion technique at a protein concentration of 17 mg/ml in a buffer consisting of 10 mM HEPES pH 7.5, 500 mM NaCL, 10% Glycerol, 300 µM ADP, 300 µM MgSO4. Crystals grew at 4°C from conditions containing 0.1M MES pH 6.0, 22% peg 20000, and were cryo protected by transferring to a cryo solution consisting of the well solution supplemented with the addition of 15% Glycerol before being plunged into liquid nitrogen.
NMR Spectroscopy:Data Collection:Data was collected at diamond light source beamline I24. Data were processed using XDS, and the structure was solved by molecular replacement using human RecQ1 (PDBid 2v1x) as a search model and the program PHASER. Model building was performed with the program COOT and the model refinement was performed using PHENIX_REFINE to a final Rfactor=20.8 % , Rfree =24.5 %.
Data Processing: