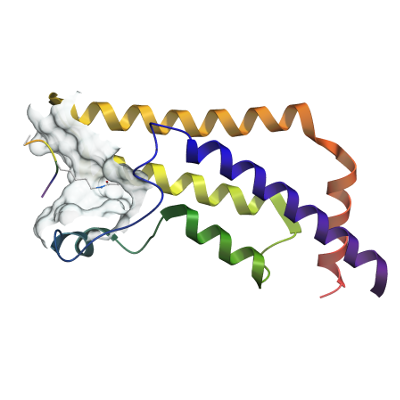
ATAD2
PDB:4QUU
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:gi|24497618
Entry Clone Source:synthetic
SGC Clone Accession:
Tag:mhhhhhhssgvdlgtenlyfq*s(m) TEV-cleavable (*) N-terminal his6 tag.
Host:BL21(DE3)-R3-pRARE2
Construct
Prelude:
Sequence:mhhhhhhssgvdlgtenlyfq*smQEEDTFRELRIFLRNVTHRLAIDKRFRVFTKPVDPDEVPDYVTVIKQPMDLSSVISKIDLHKYLTVKDYLRDIDLICSNALEYNPDRDPGDRLIRHRACALRDTAYAIIKEELDEDFEQLCEEIQESR
Vector:pNIC28-Bsa4
Growth
Medium:
Antibiotics:
Procedure:The expression plasmid was transformed into the host strain and plated on LB-agar containing 50 µg/mL kanamycin and 35 µg/mL chloramphenicol. Several colonies were combined to inoculate a 1mL culture in TB (+ 50 µg/mL kanamycin, 35 µg/mL chloramphenicol). The culture was grown overnight, glycerol was added to 15% v/v (from a 60% stock), and the resulting glycerol stock was frozen at -80 oC in 100 µL aliquots. When required a 50 mL overnight culture was started by inoculating a LB medium containing antibiotic as above. 5mL this overnight culture was used to inoculate similar 1 litre cultures (4L total) which were grown at 37oC until the OD600 reached ~0.5 at which point the temperature was adjusted to 18oC. Expression was induced overnight using 0.5 mM IPTG. The cells were collected by centrifugation and the pellet re-suspended in binding buffer and frozen. (Binding buffer: 50mM HEPES pH 7.5; 500 mM NaCl; 5 mM imidazole, 5% glycerol.)
Purification
Buffers
Procedure
Column 1: Ion exchange - Nucleic acid removal. DEAE cellulose (DE52, Whatman), 10 g of resin in 2.5 x 20 cm column. The resin was hydrated in 2.5M NaCl, washed with 20 mL binding buffer prior to loading the sample.Buffers: 50mM HEPES pH 7.5; 500 mM NaCl; 5 mM imidazoleProcedure:Supernatant was applied by gravity flow, followed by a wash with 50 mL binding buffer. The column flow Column 2: Ni-affinity. Ni-NTA (Qiagen), 5 mL of 50% slurry in 1.5 x 10 cm column, washed with binding buffer.Buffers: Binding buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 5 mM imidazole, 5% glycerol; Wash buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 30 mM Imidazole, 5% glycerol; Elution buffer: 50 mM HEPES pH 7.5, 500 mM NaCl, 50 to 250 mM Imidazole , 5% Glycerol.Procedure: The flowthrough from column 1 was loaded by gravity flow on the Ni-NTA column. The column was then washed with 100 mL wash buffer at gravity flow. The protein was eluted by gravity flow by applying 5-mL portions of elution buffer with increasing concentration of imidazole (50 mM, 100 mM, 150 mM and 250 mM); fractions were collected until essentially all protein was eluted. 10 mM DTT was added for overnight storage together with TEV protease for cleavage of the N-terminal hexahistidine tag.Column 3: Size Exclusion Chromatography. Superdex S200 16/60 HiLoadBuffers:25 mM HEPES, pH 7.5; 150 mM NaCl, 0.5 mM TCEP.Procedure: Protein was applied to a S200 gel filtration column using an ÄKTAxpress system at 1.0 mL/min. Eluted proteins were collected in 1.8-mL fractions and analyzed on SDS-PAGE. 10 mM DTT was added for storage. Column 4: Ni-affinity. Ni-NTA (Qiagen), 2 mL of 50% slurry in 1.5 x 10 cm column, washed with binding buffer. Procedure: The elution from column 3 (TEV cleaved) was loaded by gravity flow on a Ni-NTA column to remove Ni-binding contaminants. The column was washed with 10 mL wash buffer (as column 2) and proteins eluted as column 2. 10 mM DTT was added for overnight storage.
Extraction
Buffers
Procedure
Frozen pellets were thawed and fresh 1 mM PMSF and 0.5 mM TCEP were added. Cells were lysed by sonication. The lysate was centrifuged at 21,000 rpm for 60 minutes and the supernatant collected for purification.
Concentration:The cleaved purified protein was concentrated in a VivaSpin500 (5 K MWCO) to 12.2 mg/ml and stored at 4 oC. The protein concentration was determined spectrophotometrically
Ligand
MassSpec:Observed mass without histidine tag, 15430.6 (calculated mass without histidine tag, 15431).
Crystallization:Crystals were obtained using the vapor diffusion method. The protein was concentrated in gel filtration buffer to a protein concentration of 12.2 mg/mL. Sitting drops comprising 75 nl of the concentrated protein mixed with 75 nl of a well solution (2.0 M (NH4)2SO4; 0.1M BisTris pH 5.5) were equilibrated against well solution at 4°C. Crystals appeared within 2-3 days.For transfer/soaking experiments, the crystals were transferred into the stabilizing/soaking solution, containing either i) 45-50% MPD, 0.1 M bis-tris pH 5.5 and 0.1 M ammonium phosphate or ii) 28-32% PEG 3350, 50 mM bis-tris pH 5.5, 50 mM ammonium phosphate and 20% ethylene glycol. Fragments were supplemented in the soaking solution at 20-50 mM concentration (10% for NMP). Soaking time was varied from 4-9 hours.Peptide complexes: Soaking experiments were performed in a drop containing stabilizing solution (PEG 3,350, Bis-Tris, pH 5.5, ammonium phosphate) supplemented with 6 mM histone peptide and 25% ethylene glycol for 2 hours.
NMR Spectroscopy:
Data Collection:Crystals were cryo-protected using the well solution supplemented with 2M Li2SO4 and flash frozen in liquid nitrogen (apo) or using ethylene glycol present in the soaking solution . Diffraction data were collected from a single crystal on a Rigaku FR-E SuperBright at a single wavelength of 1.5 Å. The structure was solved by molecular replacement and refined to 1.89 Å.
Data Processing: