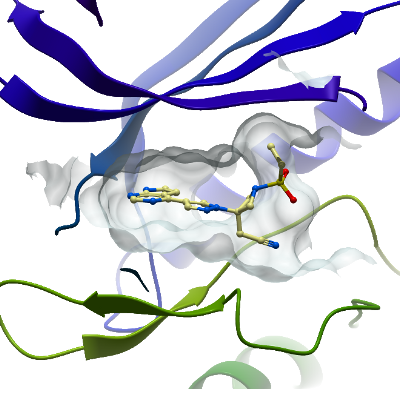
BMP2K
PDB:4W9X
Revision
Revision Type:created
Revised by:created
Revision Date:created
Entry Clone Accession:
Entry Clone Source:SGC Oxford, site-directed mutagenesis
SGC Clone Accession:
Tag:N-terminal, TEV cleavable hexahistidine tag
Host:
Construct
Prelude:
Sequence:MHHHHHHSSGVDNKFNKERRRARREIRHLPNLNREQRRAFIRSLRDDPSQSANLLAEAKKLNDAQPKGTENLYFQSMSVGVRVFAVGRHQVTLEESLAEGGFSTVFLVRTHGGIRCALKRMYVNNMPDLNVCKREITIMKELSGHKNIVGYLDCAVNSISDNVWEVLILMEYCRAGQVVNQMNKKLQTGFTEPEVLQIFCDTCEAVARLHQCKTPIIHRDLKVENILLNDGGNYVLCDFGSATNKFLNPQKDGVNVVEEEIKKYTTLSYRAPEMINLYGGKPITTKADIWALGCLLYKLCFFTLPFGESQVAICDGNFTIPDNSRYSRNIHCLIRFMLEPDPEHRPDIFQVSYFAFKFAAADCPVSNINNSSIPSALPEPMTASE
surface entropy mutations: K320A, K321A
The N-terminal residues derive from the vector:
MHHHHHHSSGVDNKFNKERRRARREIRHLPNLNREQRRAFIRSLRDDPSQSANLLAEAKKLNDAQPKGTENLYFQS, derive from the vector.
Vector:pNIC-ZB
Growth
Medium:
Antibiotics:
Procedure:
Purification
Buffers
Procedure
Cell Lysis: The resuspended cell pellet was thawed and lysed by sonication. PEI (polyethyleneimine) was added to a final concentration of 0.15 %. The cell debris and precipitated DNA were spun down at 22,000 rpm for 1 hour, 4°C.Lysis Buffer:50 mM Hepes pH 7.5, 500 mM NaCl, 5 mM Imidazole, 5 % glycerol, 0.5 mM TCEP, 1:2000 dilution protease inhibitor cocktail. Purification: Column 1: 2.5 ml (final bed volume) of Ni-Sepharose in a 2 cm diameter gravity flow column, equilibrated in binding buffer. Column 1 Buffers: Binding Buffer:50 mM Hepes pH 7.5, 500 mM NaCl, 5 mM Imidazole, 5 % glycerol, 0.5 mM TCEP. Wash Buffer:As Binding Buffer except 30 mM imidazole. Elution Buffers 1-4: As Binding Buffer except 50, 100, 150 or 250 mM imidazole. Column 1 Procedure:The clarified supernatant was filtered through a 0.45 uM pore size syringe filter and passed through the column. The column was washed with 50 ml of Binding Buffer and 30 ml of Wash Buffer. 5 ml of each Elution Buffer was passed through to elute the protein in a stepwise manner, finally a second 5 mL of Elution Buffer 4 was added. Tag Cleavage: Tacacco etch virus (TEV) protease added (200 µL at 2 mg/mL) to the pooled elution fractions and incubated at 4°C overnight. Column 2: S200 16/60 Gel Filtration (GE Healthcare) Column 2 Buffers: GF Buffer:50 mM Hepes pH 7.5, 300 mM NaCl, 5 % glycerol, 0.5 mM TCEP, Column 2 Procedure:The eluted protein was concentrated to 4 ml volume and injected onto the column. Column 3: 1 mL Ni-Sepharose in gravity flow column, pre-equilibrated in binding buffer. Column 3 Buffers: As per Column 1 Column 3 Procedure: Pooled fractions from gel filtration were added to the column and the flow-through collected. 3 mL of wash buffer added, followed by 3 mL of elution buffer 4 (250 mM imidazole) and fractions pooled separately.
Extraction
Buffers
Procedure
Expression strain: BL-21(DE3)-R3-lambda-ppase (A homemade phage resistant version of BL21(DE3) that carries a plasmid for co-expression of lambda phosphatase). Transformation:The construct DNA was transformed into homemade chemically competent cells of the expression strain by a standard heat shock procedure. Expression:Colonies were used to inoculate 10 ml of LB media containing 50 µg/ml kanamycin and 34 µg/ml chloramphenicol in a 50 ml falcon tube, which was placed in a 37°C shaker overnight. The next day 3x 3 ml of this starter culture was used to inoculate 3x 1L of LB media containing 35 µg/ml kanamycin in 2L baffled shaker flasks. When the OD600 was approximately 0.45, the temperature was reduced to 20°C and when the OD600 was approximately 0.6 the cells were induced by the addition of 0.5 mM IPTG. The expression was continued overnight. Cell harvest:Cells were spun at 5000rpm for 15 mins and the pellets resuspended in Lysis Buffer and then frozen at -20°C.
Concentration:The BMP2K protein flow-through from Column 3 was concentrated to 13 mg/ml (measured by 280 nm absorbance).
Ligand
MassSpec:Expected: 34754.1Observed: 34754.2
Crystallization:Complex preparation clinical kinase inhibitor compounds:Protein spun at high speed (14,000 rpm for 10 minutes at 4 degrees) to remove any aggregates. Clinical kinase inhibitors: AZD7762 and BaricitinibFrom 50 mM compound stocks in DMSO, 2 µL compound added to 65 µL protein and incubated on ice for 10-20 minutes prior to setting up plates. (1.5 mM final compound concentration + 3 % DMSO). Crystallisation: 3-drop ratios (2:1, 1:1, 1:2 of protein to reservoir well solution) in 150 nL total volume equilibrated against 20 µL reservoir well solution in 96-well sitting-drop vapour diffusion plates. Crystals grew in a range of conditions at 293 K.further optimisation of conditions was required including addition of a seed stock solution generated using crushed crystals from previous rounds of optimisation. Best diffracting crystals grew from 4.5M NaCl, 0.1M bis-Tris pH 5.5.
NMR Spectroscopy:
Data Collection:Crystals were cryo-protected in a solution containing mother liquor supplemented with 25 % ethylene glycol, then flash-frozen in liquid nitrogen. Data was collected at Diamond beamline I04-1.
Data Processing: