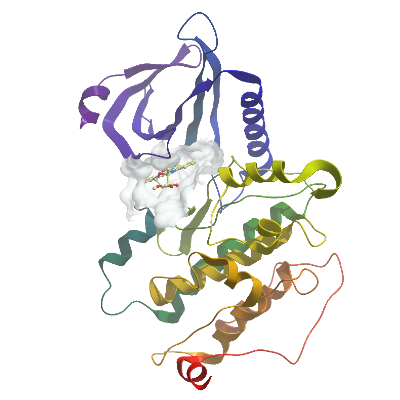
No. XX AAK1A PDB: 4WSQ Materials & Methods |
Entry Clone Source: MGC |
Entry Clone Accession: |
SGC Construct ID: AAK1A-c046 |
Coding DNA sequence:
ATGACCTCGGGCCTGGGCAGTGGCTA
CATCGGAAGAGTCTTCGGCATCGGGC
GACAGCAGGTCACAGTGGACGAGGTG
TTGGCGGAAGGTGGATTTGCTATTGT
ATTTCTGGTGAGGACAAGCAATGGGA
TGAAATGTGCCTTGAAACGCATGTTT
GTCAACAATGAGCATGATCTCCAGGT
GTGCAAGAGAGAAATCCAGATAATGA
GGGATCTTTCAGGGCACAAGAATATT
GTGGGTTACATTGATTCTAGTATCAA
CAACGTGAGTAGCGGTGATGTATGGG
AAGTGCTCATTCTGATGGACTTTTGT
AGAGGTGGCCAGGTGGTAAACCTGAT
GAACCAGCGCCTGCAAACAGGCTTTA
CAGAGAATGAAGTGCTCCAGATATTT
TGTGATACCTGTGAAGCTGTTGCCCG
CCTGCATCAGTGCAAAACTCCTATTA
TCCACCGGGACCTGAAGGTTGAAAAC
ATCCTCTTGCATGACCGAGGCCACTA
TGTCCTGTGTGACTTTGGAAGCGCCA
CCAACAAATTCCAGAATCCACAAACT
GAGGGAGTCAATGCAGTAGAAGATGA
GATTAAGAAATACACAACGCTGTCCT
ATCGAGCACCAGAAATGGTCAACCTG
TACAGTGGCAAAATCATCACTACGAA
GGCAGACATTTGGGCTCTTGGATGTT
TGTTGTATAAATTATGCTACTTCACT
TTGCCATTTGGGGAAAGTCAGGTGGC
AATTTGTGATGGAAACTTCACAATTC
CTGATAATTCTCGATATTCTCAAGAC
ATGCACTGCCTAATTAGGTATATGTT
GGAACCAGACCCTGACAAAAGGCCGG
ATATTTACCAGGTGTCCTACTTCTCA
TTTAAGCTACTCAAGAAAGAGTGCCC
AATTCCAAATGTACAGAACTCTCCCA
TTCCTGCAAAGCTTCCTGAACCAGTG
AAAGCCAGTGAGGCAGCTGCAAAAAA
GACCCAGCCAAAGGCCAGACTGACAG
ATCCCATTCCCACCACAGAGACTTCA
ATTGCAGCAGAGAACCTCTACTTCCA
ATCGCACCATCATCACCACCATTGA
|
Expressed protein sequence:
MTSGLGSGYIGRVFGIGRQQVTVDEV
LAEGGFAIVFLVRTSNGMKCALKRMF
VNNEHDLQVCKREIQIMRDLSGHKNI
VGYIDSSINNVSSGDVWEVLILMDFC
RGGQVVNLMNQRLQTGFTENEVLQIF
CDTCEAVARLHQCKTPIIHRDLKVEN
ILLHDRGHYVLCDFGSATNKFQNPQT
EGVNAVEDEIKKYTTLSYRAPEMVNL
YSGKIITTKADIWALGCLLYKLCYFT
LPFGESQVAICDGNFTIPDNSRYSQD
MHCLIRYMLEPDPDKRPDIYQVSYFS
FKLLKKECPIPNVQNSPIPAKLPEPV
KASEAAAKKTQPKARLTDPIPTTETS
IAAENLYFQSHHHHHH
|
Vector: pNIC-CTH0
C-terminal His, TEV-cleavable, for LIC cloning.
|
Tags and additions: AENLYFQSHHHHHH: C-terminal hexahistidine tag cleavable by TEV protease. |
Host: BL21 (DE3)-R3-pRARE2. Phage-resistant derivative of BL21 (DE3), with pRARE2 plasmid encoding rare codon tRNAs (chloramphenicol-resistant). |
Growth Medium & Induction Protocol: A glycerol stock was used to inoculate a 10ml starter culture containing LB media with 50µg/ml Kanamycin + 34 µg/ml Chloramphenicol . The starter culture was grown overnight at 37°C with shaking at 200 rpm. The following morning, flasks containing 1L TB/Kanamycin were each inoculated with 5 ml of the starter culture. Cultures were incubated at 37°C with shaking at 170 rpm until an OD600nm >= 2.0 was reached. The flasks were then cooled down to 18°C and 0.4mM IPTG added to induce protein expression overnight. Cells were harvested by centrifugation at 5000 rpm at 4°C for 15 min. Cell pellets from each flask were resuspended in 15ml Binding buffer (50mM HEPES, pH 7.5; 500mM NaCl; 5% Glycerol; 5mM Imidazole; 0.5mM TCEP; 1:2000 Protease Inhibitor Cocktail) and frozen at -20°C. |
Extraction buffer, extraction method: The frozen cells were thawed. The cells were lysed by ultrasonication over 15 min with the sonicator pulsing ON for 5 sec and OFF for 10. A final concentration of 0.15% PEI was added to the lysate. The cell lysate was spun down by centrifugation at 21.5K rpm at 4°C for 1 h. The supernatant was recovered for purification.
|
Column 1: Ni-Affinity Chromatography. 5 ml of 50 % Ni-sepharose slurry was applied onto a 1.5 x 10 cm column. The column was equilibrated with binding buffer (50ml). |
Buffers:
Binding buffer: 50 mM Hepes, pH 7.5; 500 mM NaCl; 5% Glycerol; 5 mM imidazole 0.5mM TCEP
Wash buffer: 50 mM Hepes, pH 7.5; 500 mM NaCl; 5% Glycerol; 30 mM imidazole 0.5mM TCEP
Elution buffer: 50 mM HEPES, pH 7.5; 500 mM NaCl; 5% Glycerol; 50, 100, 150 and 250 mM imidazole 0.5mM TCEP
|
Procedure: The supernatant following centrifugation was applied by gravity flow onto the Ni-sepharose column. The bound protein was then washed with 50ml binding buffer and subsequently with 30 ml wash buffer. AAK1A protein was then eluted by applying a step gradient of imidazole - using 5 ml portions of elution buffer with increasing concentration of imidazole (1 x 50 mM, 1 x 100 mM, 1 x 150 mM and 2 x 250 mM). Fractions were analyzed by SDS PAGE and the second, third, fourth and fifth elution fractions were kept and pooled. |
Enzymatic treatment: TEV protease cleavage. Fractions containing AAK1A were treated with TEV protease overnight at 4°C. |
Column 2: Size Exclusion Chromatography - S200 HiLoad 16/60 Superdex run on ÄKTA-Express |
Gel Filtration buffer: 300 mM NaCl, 50 mM HEPES pH 7.5, 0.5 mM TCEP, pH 7.5, 5% glycerol
|
Procedure: The Superdex S200 column was first equilibrated with Gel Filtration buffer. The protein fraction from above step was concentrated to |
Column 3: Size Exclusion Chromatography - S200 HiLoad 16/60 Superdex run on ÄKTA-Express Cation Exchange Chromatography. HiTrap SP HP 5ml column on ÄKTA-Express |
QA Buffer I: 50mM HEPES pH 7.5
QB Buffer II: 1M NaCl, 50 mM HEPES pH 7.5
|
Procedure: The HiTrap Q HP column was equilibrated with 50mL QA buffer I. The protein fraction from above step was concentrated to 5mL using a centrifugal filter with a 10kDa cut-off. The 5mL fraction was made up to 50ml with QA buffer I and applied onto the column. Bound protein was eluted in 0%-100% gradient with QB buffer II. Several peaks were found corresponding to different phosphorylation states of the protein. Peaks were pooled separately. |
Crystallization: Dephosphorylated protein was concentrated to 11 mg/mL. The compound K252a was added to a final concentration of 1.5 mM from a stock at 50 mM in 100% DMSO. Prior to setting up crystallization plates the solution of AAK1-K252a was incubated on ice for approximately 30 minutes. Sitting-drop vapour diffusion plates were prepared. Crystals grew under multiple conditions using freshly prepared protein. The best-diffracting crystals of the AAK1-K252a complex were obtained by mixing 50 nL of protein with 100 nL of a reservoir solution containing 20% PEG 3350, 0.5 mM Zinc Acetate. Prior to mounting crystals were cryo-protected in situ by addition of reservoir solution containing an additional 25% ethylene glycol. Crystals were then flash frozen in liquid nitrogen. |
Data Collection:2.1 Å resolutionX-ray source: Diamond Light Source, station I04, using monochromatic radiation at wavelength 0.92 Å
Crystals belonged to spacegroup P212121 with unit cell parameters a=68.66 Å b=71.32 Å c=183.64 Å, α=90° β= 90 γ= 90°. Diffraction data were integrated and scaled using Mofslm and SCALA, respectively. Molecular replacement was used in Phaser MR and PDB entry 2buj was used as a search model. Two molecules were present in the asymmetric unit. COOT, REFMAC and PHENIX.REFINE were used for model building and refinement..
|