Molecular Biology
Entry Clone Accession: MGC:22457 IMAGE:4767736
Entry Clone Source: MGC
SGC Construct ID: PAHA-c011
Protein Region: G19-V118
Vector: pNIC28-Bsa4
Tag: N-6HIS; TEV cleavage site
Host: BL21(DE3)-R3-pRARE2
Sequence (with tag(s)): MHHHHHHSSGVDLGTENLYFQSMGQETSYIEDNCNQNGAISLIFSLKEEVGALAKVLRLFEENDVNLTHIESRPSRLKKDEYEFFTHLDKRSLPALTNIIKILRHDIGATVHELSRDKKKDTV
Sequence after tag cleavage: SMGQETSYIEDNCNQNGAISLIFSLKEEVGALAKVLRLFEENDVNLTHIESRPSRLKKDEYEFFTHLDKRSLPALTNIIKILRHDIGATVHELSRDKKKDTV
DNA sequence:
CATATGCACCATCATCATCATCATTCTTCTGGTGTAGATCTGGGTACCGAGAACCTGTACTTCCAATCCATGGGACAGGAAACAAGCTATATTGAAGACAACTGCAATCAAAATGGTGCCATATCGCTGATCTTCTCACTCAAAGAAGAAGTTGGTGCATTGGCCAAAGTATTGCGCTTATTTGAGGAGAATGATGTAAACCTGACCCACATTGAATCTAGACCTTCTCGTTTAAAGAAAGATGAGTATGAATTTTTCACCCATTTGGATAAACGTAGCCTGCCTGCTCTGACAAACATCATCAAGATCTTGAGGCATGACATTGGTGCCACTGTCCATGAGCTTTCACGAGATAAGAAGAAAGACACAGTGTGACAGTAAAGGTGGATACGGATCCGAA
Protein Expression
Medium: Terrific Broth
Antibiotics: Kanamycin
Procedure: plasmid was transformed into E. coli BL21(DE3), cultured in Terrific Broth at 37oC until OD600 ~1.5, and induced with 0.5 mM IPTG for overnight growth at 18°C.
Protein Purification
Procedure: Cells were harvested and homogenized in buffer A (50 mM HEPES pH 7.5, 500 mM NaCl, 5 mM imidazole, 0.5 mM TCEP, EDTA-free protease inhibitor). Insoluble debris was removed by further centrifugation. Proteins were purified by passing cell extracts through a 1ml HisTrap column pre-equilibrated with buffer A, and eluted with buffer B (buffer A + 250 mM Imidazole). Eluted fractions were treated with TEV protease at a protein:protease ratio of 1:20, and incubated overnight at 4°C in order to cleave off the His6-tag. The tag-cleaved protein was applied onto a 1ml HisTrap column, pre-equilibrated with GF buffer (50 mM HEPES, pH 7.5, 300 mM NaCl, 5% glycerol and 0.5 mM TCEP). The flow-through sample was applied onto a HiLoad 16/60 Superdex 75 column pre-equilibrated with GF buffer.
Concentration: 13.5 mg/ml
Mass-spec Verification: Yes
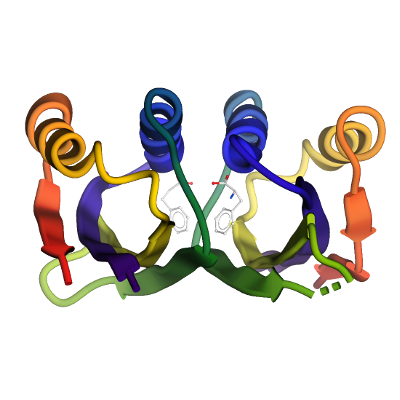
Structure Determination
Crystallization: Protein was concentrated to a 13.5mg/ml, where l-Phenylalanine (Phe) was added to a final concentration of 10mM. The sample was treated with trypsin at a protein:trypsin mass ratio of 1:100 immediately prior to crystallization set up. Crystals were grown by vapour diffusion in sitting drops at 20°C. A sitting drop consisting of 100 nl trypsin-treated protein and 200 nl well solution was equilibrated against well solution containing 25% PEG 3350, 0.20 M NaCl and 0.1 M BIS-Tris pH 5.5. The crystals were mounted directly from the drop using 25% ethylene glycol as a cryoprotectant and flash-cooled in liquid nitrogen.
Data Collection: Diffraction data was collected at the Diamond Light Source beamline Dmnd I03; Resolution: 1.8 Å
Data Processing: Diffraction data were processed with the CCP4 program suite. The structure was solved by molecular replacement using the program PHASER with the Chlorobium tepidum prephenate dehydratase structure (PDB code 2QMX) as search model. Iterative cycles of restrained refinement and manual model building were performed using COOT and PHENIX.