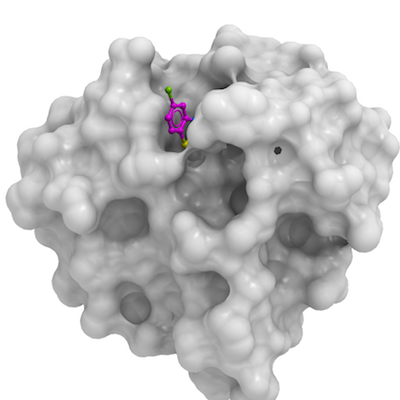
HDAC6
PDB:5KH3
Entry Clone Accession:BC013737
Entry Clone Source:MGC AT36-C10
SGC Clone Accession:JMC105-C01
Tag:N-terminal His6-tag, removed
Host:BL21(DE3)V2R-pRARE2
Vector:pET28-lic
Prelude:HDAC6:P1109-H1213. The sequence is consistent with reference sequence NP_006035.
Sequence: gsPLPWCPHLVAVCPIPAAGLDVTQPCGDCGTIQENWVCLSCYQVYCGRYINGHMLQHHGNSGHPLVLSYIDLSAWCYYCQAYVHHQALLDVKNIAHQNKFGEDMPH
Growth
Procedure: Shaker. Cultures were grown in M9 media supplemented with 50 µM ZnCl2, and grown at 37 °C until OD600 reached 0.6 before induction with 0.2 mM IPTG and overnight growth at 15 °C.
Purification
Buffers: Washing Buffer: 50 mM tris-pH8, 300 mM NaCl, 25 mM imidazole, 2 mM TCEP - Elution Buffer: 50 mM tris-pH8, 300 mM NaCl, 300 mM imidazole, 2 mM TCEP - Dialysis Buffer: 50 mM tris-pH8, 150 mM NaCl, 2 mM TCEP, 2 mM CaCl2 - Size-exclusion buffer: 50 mM tris-pH8, 150 mM NaCl, 2 mM TCEP
Procedure: Following sonication, lysates were clarified by centrifugation, 15000 rpm, 1 hour, JLA16.250. Proteins were purified first using Ni-affinity chromatography with NiNTA resin (Qiagen) (1 mL slurry per L cells) washing 100 CV extraction buffer then 100 CV washing buffer and then eluted with 5 CV elution buffer. The eluted protein was dialysed against 50 mM tris-pH8, 150 mM NaCl, 2 mM TCEP overnight (snakeskin MWCO 3500) and the his-tags were cleaved with thrombin for protein. Samples were then concentrated (Amicon MWCO 10,000) and further purified by gel filtration using Superdex 75 16/60 before being concentrated to 5 mg/ml.
Extraction
Buffers: 50 mM tris-pH8, 300 mM NaCl, 5 mM imidazole, 2 mM TCEP
Procedure: Cell pastes were resuspended in 20 fold volumes of lysis buffer (50 mM tris-pH8, 300 mM NaCl, 5 mM imidazole, 2 mM TCEP) supplemented with benzonase and protease inhibitors. Cell suspensions were sonicated by 10 mins at 5" on 7" off duty cycle at 120W output power.
Concentration: Concentration used for crystallization : native protein: 3.5 mg/mL
Ligand: Zn, 3-(5-Chloro-1,3-benzothiazol-2-yl)propanoic acid
Structure Determination
MassSpec:11920.62 g/mol
Crystallization:The apo crystal structure of the HDAC6 ZnF-UBD 1109-1215 was previously solved at SGC Toronto (PDB ID: 3C5K). These crystals can be obtained by mixing the protein solution at 3.5 mg/ml 1:1 with 3.5M sodium formate, 0.1M bis tris propane, 5% ethylene glycol, pH7 using the vapour diffusion method at room temperature. In this crystal form, the primary binding pocket is occluded by the C-terminus of the adjacent protein molecule in the crystal lattice, although a secondary adjacent site is exposed to the solvent. These crystals can be used to seed for the desired crystal form for fragment screening. Diluting a 1 µl drop containing these crystals 1:10,000 with mother liquor and vortexing the sample vigourously yields a seed mix. HDAC6 ZF UBD 1109-1213 can then be crystallised in a high salt condition containing 2M Na formate, 0.1M Na acetate pH4.6, 5% ethylene glycol, again by the vapour diffusion method. 500nl of protein are added to 400nl mother liquor and then 100nl seed mix per drop, typically plates are set up using a mosquito. These crystals have a solvent exposed ubiquitin binding pocket, amenable to soaking with 5 % DMSO-compound stock for 3 hours and can be cryo cooled in liquid nitrogen without additional cryo-protection.