Molecular Biology
Entry Clone Accession: clone 27 (T615S mutation)
Entry Clone Source: Collaborator
SGC Construct ID: AASSA-c013
Protein Region: L455-P926
Vector: pFB-LIC-Bse
Tag: N-6HIS;N-TEV
Host: DH10Bac
Sequence (with tag(s)): MGHHHHHHSSGVDLGTENLYFQSMALPDKYKYIQTLRESRERAQSLSMGTRRKVLVLGSGYISEPVLEYLSRDGNIEITVGSDMKNQIEQLGKKYNINPVSMDICKQEEKLGFLVAKQDLVISLLPYVLHPLVAKACITNKVNMVTASYITPALKELEKSVEDAGITIIGELGLDPGLDHMLAMESIDKAKEVGATIESYISYCGGLPAPEHSNNPLRYKFSWSPVGVLMNVMQSATYLLDGKVVNVAGGISFLDAVTSMDFFPGLNLEGYPNRDSTKYAEIYGISSAHTLLRGTLRYKGYMKALNGFVKLGLINREALPAFRPEANPLTWKQLLCDLVGISPSSEHDVLKEAVLKKLGGDNTQLEAAEWLGLLGDEQVPQAESILDALSKHLVMKLSYGPEEKDMIVMRDSFGIRHPSGHLEHKTIDLVAYGDINGFSAMAKTVGLPTAMAAKMLLDGEIGAKGLMGPFSKEIYGPILERIKAEGIIYTTQSTIKP
Sequence after tag cleavage: SMALPDKYKYIQTLRESRERAQSLSMGTRRKVLVLGSGYISEPVLEYLSRDGNIEITVGSDMKNQIEQLGKKYNINPVSMDICKQEEKLGFLVAKQDLVISLLPYVLHPLVAKACITNKVNMVTASYITPALKELEKSVEDAGITIIGELGLDPGLDHMLAMESIDKAKEVGATIESYISYCGGLPAPEHSNNPLRYKFSWSPVGVLMNVMQSATYLLDGKVVNVAGGISFLDAVTSMDFFPGLNLEGYPNRDSTKYAEIYGISSAHTLLRGTLRYKGYMKALNGFVKLGLINREALPAFRPEANPLTWKQLLCDLVGISPSSEHDVLKEAVLKKLGGDNTQLEAAEWLGLLGDEQVPQAESILDALSKHLVMKLSYGPEEKDMIVMRDSFGIRHPSGHLEHKTIDLVAYGDINGFSAMAKTVGLPTAMAAKMLLDGEIGAKGLMGPFSKEIYGPILERIKAEGIIYTTQSTIKP
DNA Sequence: CCATGGGCCACCATCATCATCATCATTCTTCTGGTGTAGATCTGGGTACCGAGAACCTGTACTTCCAATCCATGGCATTACCTGATAAATATAAATATATCCAGACACTCCGGGAGAGCAGGGAACGTGCTCAGTCACTTTCAATGGGCACCAGGAGAAAGGTTTTGGTTCTTGGATCTGGCTACATATCTGAGCCTGTATTAGAATATTTATCAAGAGATGGCAATATAGAAATAACAGTAGGATCTGACATGAAGAATCAAATTGAACAGTTAGGCAAGAAATATAATATTAATCCTGTTAGCATGGACATTTGTAAACAAGAAGAGAAGCTGGGCTTCTTGGTGGCAAAACAGGATCTTGTCATCAGCTTGTTGCCTTATGTATTGCACCCTCTTGTGGCCAAGGCCTGCATCACAAACAAAGTTAACATGGTCACTGCAAGCTACATCACACCAGCACTAAAAGAATTGGAAAAGAGTGTGGAAGATGCTGGCATCACAATCATTGGTGAATTGGGATTGGACCCTGGTCTGGATCACATGTTAGCAATGGAATCAATAGATAAAGCCAAGGAAGTGGGAGCCACGATTGAATCATATATTTCCTACTGTGGTGGGCTTCCAGCCCCTGAACATTCAAACAATCCATTGAGATATAAATTTAGCTGGAGTCCAGTGGGAGTTTTGATGAATGTAATGCAGTCTGCCACCTATCTGCTCGATGGAAAGGTTGTGAATGTTGCAGGAGGCATCTCCTTTCTTGATGCCGTTACGTCCATGGATTTTTTTCCAGGATTAAATTTGGAAGGCTATCCTAACAGAGACAGTACGAAATATGCTGAGATTTATGGCATTTCTTCTGCTCACACTTTGTTGCGGGGGACACTGAGATATAAGGGATATATGAAAGCTTTGAATGGATTTGTAAAATTAGGTCTTATAAACAGAGAAGCGCTTCCTGCCTTTAGACCTGAGGCCAACCCTCTCACCTGGAAACAACTCCTCTGTGACCTAGTTGGGATTTCACCCTCCTCTGAGCATGATGTGTTGAAGGAAGCTGTTCTTAAGAAACTAGGAGGAGACAATACCCAGTTGGAGGCTGCTGAATGGTTGGGCTTACTTGGGGATGAACAAGTTCCTCAGGCAGAGTCCATTCTGGATGCCCTCTCCAAGCATTTGGTCATGAAGCTTTCCTATGGTCCTGAAGAAAAAGATATGATTGTGATGAGAGACAGCTTTGGAATCAGACATCCTTCTGGACATTTAGAACATAAAACGATTGATCTTGTGGCTTATGGGGACATCAATGGCTTTTCAGCCATGGCTAAAACCGTGGGGTTACCCACCGCCATGGCAGCCAAAATGTTGCTTGATGGTGAAATTGGAGCCAAAGGCCTAATGGGGCCCTTTTCAAAGGAGATCTATGGACCAATATTGGAGCGAATTAAAGCAGAAGGCATTATATATACTACACAGAGTACAATTAAACCATGACAGTAAAGGTGGATACGGATCCGAATTCGAGCTCCGTCGACAAGCTT
Protein Expression
Medium: InsectXpress medium (Lonza)
Antibiotics: Ampicillin
Procedure: Bacmid DNA was prepared from DH10Bac cells and using to transfect Sf9 insect cells for the preparation of initial baculovirus. AASS protein was expressed from infected Sf9 cells cultivated in InsectXpress medium (Lonza) for 72 hours at 27°C.
Protein Purification
Buffers:
Binding Buffer: 50 mM HEPES pH 7.4, 500 mM NaCl, 5% Glycerol, 20 mM Imidazole pH 7.4, 0.5 mM TCEP
Wash Buffer: 50 mM HEPES pH 7.4, 500 mM NaCl, 5% Glycerol, 40 mM Imidazole pH 7.4, 0.5 mM TCEP
Elution Buffer: 50 mM HEPES pH 7.4, 500 mM NaCl, 5% Glycerol, 250 mM Imidazole pH 7.4, 0.5 mM TCEP
Procedure:
Harvested cells were resuspended in lysis buffer (50 mM HEPES pH 7.4, 500 mM NaCl, 5% Glycerol, 20 mM Imidazole pH 7.4, 0.5 mM TCEP, 1 µL per 1 mL protease inhibitor cocktail EDTA-free).
Cell pellet was dissolved in approximately 200 mL lysis buffer and broken by homogenization by 2 passes at 12,000 psi. The cell debris was pelleted at 35000 x g, 1h and the supernatant used for purification on a gravity flow Ni-NTA column (5 mL).
The clarified cell extract was added to 5 ml of Ni-NTA resin pre-equilibrated with lysis buffer and passed through a glass column. The column was then washed with Binding Buffer (2 x 50 mL) and Wash Buffer (2 x 50 mL). The protein was eluted with Elution Buffer in 5 x 5 mL fractions. The eluted fractions from column 1 were pooled and concentrated to 5 mL with a 30 kDa MWCO spin concentrator and injected into an S200 16/60 column (pre-equilibrated in GF Buffer (50 mM HEPES pH 7.4, 500 mM NaCl, 0.5 mM TCEP, 5% Glycerol)) at 1.0 mL/min. 1.5 mL-fractions were collected. The eluted protein was cleaved overnight at 4 °C by TEV protease (1/20 (w/w)).
The following day protein sample was loaded onto 0.5ml Ni-sepharose column pre-equilibrated with GF buffer to remove uncleaved protein. Pooled protein fractions were concentrated to 13 mg/mL using a 30 kDa mwco concentrator.
Concentration: 18.0 mg/ml
Mass-spec Verification: yes
Structure Determination
Crystallization: Apo crystals were prepared by mixing 50 nL of hAASS-SDH protein (80 mg/mL) with 100 nL of reservoir solution containing 20% PEG3350, 0.1M Tris pH 7.5 and 0.2-0.33 M sodium malonate.
Data Collection: Beamline: Dmnd I04-1; Resolution: 2.6 Å
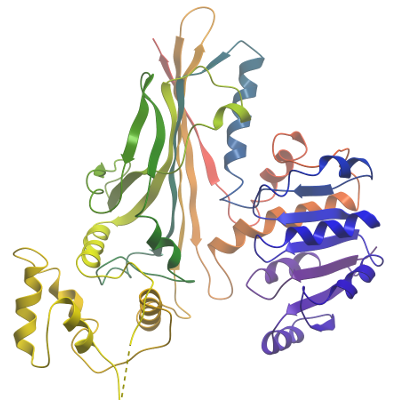
Data Processing: The structure of hAASS-SDH was solved by molecular replacement with the program PHASER, using the fungal saccharopine reductase from S.cerevisiae (PDB code 2AXQ) as search model (38% sequence identity). Two molecules were found in the asymmetric unit (a.u.). The initial model was rebuilt using phenix.autobuild. To complete the model iterative cycles of phenix.refine including TLS refinement followed by manual model building for missing residues using coot were performed. No NCS restraints were applied due to conformational differences in domain III (res 278-376) of the two copies in the a.u. Solvent atoms were placed during the last four rounds of refinement using phenix.refine.