Molecular Biology
Entry Clone Accession: IMAGE:6149862
Entry Clone Source: MGC
SGC Construct ID: BRD7A-c005
Protein Region: E129-S239
Vector: pNIC28-Bsa4
Tag: N-terminus His6 tag followed by a TEV cleavage site sequence
Host: BL21(DE3)-R3-pRARE2
Sequence (with tag(s)): MHHHHHHSSGVDLGTENLYFQSMEEVEQTPLQEALNQLMRQLQRKDPSAFFSFPVTDFIAPGYSMIIKHPMDFSTMKEKIKNNDYQSIEELKDNFKLMCTNAMIYNKPETIYYKAAKKLLHSGMKILSQERIQS
Sequence after tag cleavage: SMEEVEQTPLQEALNQLMRQLQRKDPSAFFSFPVTDFIAPGYSMIIKHPMDFSTMKEKIKNNDYQSIEELKDNFKLMCTNAMIYNKPETIYYKAAKKLLHSGMKILSQERIQS
DNA Sequence: CATATGCACCATCATCATCATCATTCTTCTGGTGTAGATCTGGGTACCGAGAACCTGTACTTCCAATCCATGGAAGAAGTAGAACAGACACCCCTTCAAGAAGCTTTGAATCAACTGATGAGACAATTGCAGAGAAAAGATCCAAGTGCTTTCTTTTCATTTCCTGTGACTGATTTTATTGCTCCTGGCTACTCCATGATCATTAAACACCCAATGGATTTTAGTACCATGAAAGAAAAGATCAAGAACAATGACTATCAGTCCATAGAAGAACTAAAGGATAACTTCAAACTAATGTGTACTAATGCCATGATTTACAATAAACCAGAGACCATTTATTATAAAGCTGCAAAGAAGCTGTTGCACTCAGGAATGAAAATTCTTAGCCAGGAAAGAATTCAGAGCTGACAGTAAAGGTGGATACGGATCCGAA
Protein Expression
Medium: TB media complemented with 50 µg/mL of kanamycin and 34 µg/mL of chloramphenicol.
Antibiotics: Kanamycin and chloramphenicol.
Procedure: Each 1L culture was inoculated with 10 mL of a starter culture grown at 37°C overnight. Then, expression was induced with 0.15 mM IPTG when the OD600nm reached 1.2, and incubated at 18°C overnight.
Protein Purification
Procedure: After expression, cultures were centrifuged at 3,500 g and pellets were frozen with liquid nitrogen. 10 g of bacterial pellet were resuspended in 100 mL of buffer A (50 mM HEPES pH 7.5, 500 mM NaCl, 10 mM imidazole, 5% glycerol, 0.5 mM TCEP) containing cOmplete EDTA-free protease inhibitors from Roche. Cell disruption was carried out using an Avestin EmulsiFlex-c5 homogenizer. Lysates were centrifuged at 35,000 g for 1h at 4°C. Samples were incubated with 2 mL of Ni-Sepharose 6 FF 1h at 4°C. After removing the non-bound lysate, beads were washed with buffer A followed by an extra wash with buffer A with 50 mM imidazole. Elution was achieved with buffer A containing 150 mM imidazole. Samples were then dialyzed into buffer A at 4°C overnight, in the presence of TEV protease. To keep the untagged BRD7, a second affinity chromatography using Ni-Sepharose 6 FF was performed. The flow through was then concentrated with an Amicon 3K centrifugal filter, and applied to a size exclusion chromatography column HiLoad 16/600 superdex 75 pg in a buffer containing 10 mM HEPES pH 7.5, 500 mM NaCl, 5% glycerol and 0.5 mM TCEP. The peak corresponding with BRD7 was concentrated to 4.24 mg/mL. Purity and protein identity were confirmed using SDS-PAGE, Q-TOF mass spectrometry, and tryptic digestion. The final yield was 2.2 mg of protein per 1 L of culture.
Columns: Column 1: 2 mL Ni-NTA; Column 2: 2 mL Ni-NTA; Column 3: HiLoad 16/600 Superdex 75
Concentration: 4.24 mg/ml
Mass-spec Verification: Yes
Structure Determination
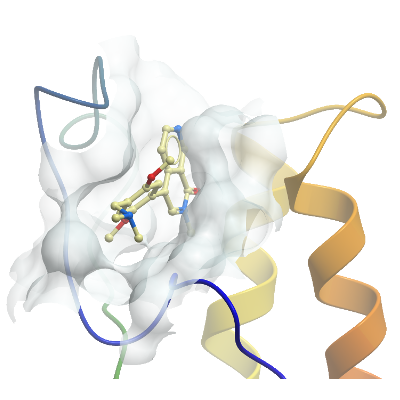
Crystallization: Prior crystallization, BRD7 was incubated with the compound BI9864 (1:3 protein:compound molar ratio) for 1h at 4°C. Crystal plates were incubated at 4°C for 5 days. Crystals grew in a condition containing 20% PEG6000, 10% ethylene glycol, 0.1 M HEPES pH 7.0, 0.01 M ZnCl2. Crystals were cryo-protected with 20% ethylene glycol, and frozen in liquid nitrogen.
Data Collection: Diffraction data were collected at Diamond Light Source beamline I04-1 using a Pilatus 6M-F detector.
Data Processing: Data were indexed and integrated using XDSgui, and scaled and analyzed with AIMLESS from the CCP4i suite. The structure was solved using Phaser. The search model was an internal BRD9 structure, which shares a 37.6% sequence homology with BRD7. The resolution obtained was 1.5 Å. Refinement was carried out using REFMAC, and restrains for BI9864 were obtained with PRODUG.