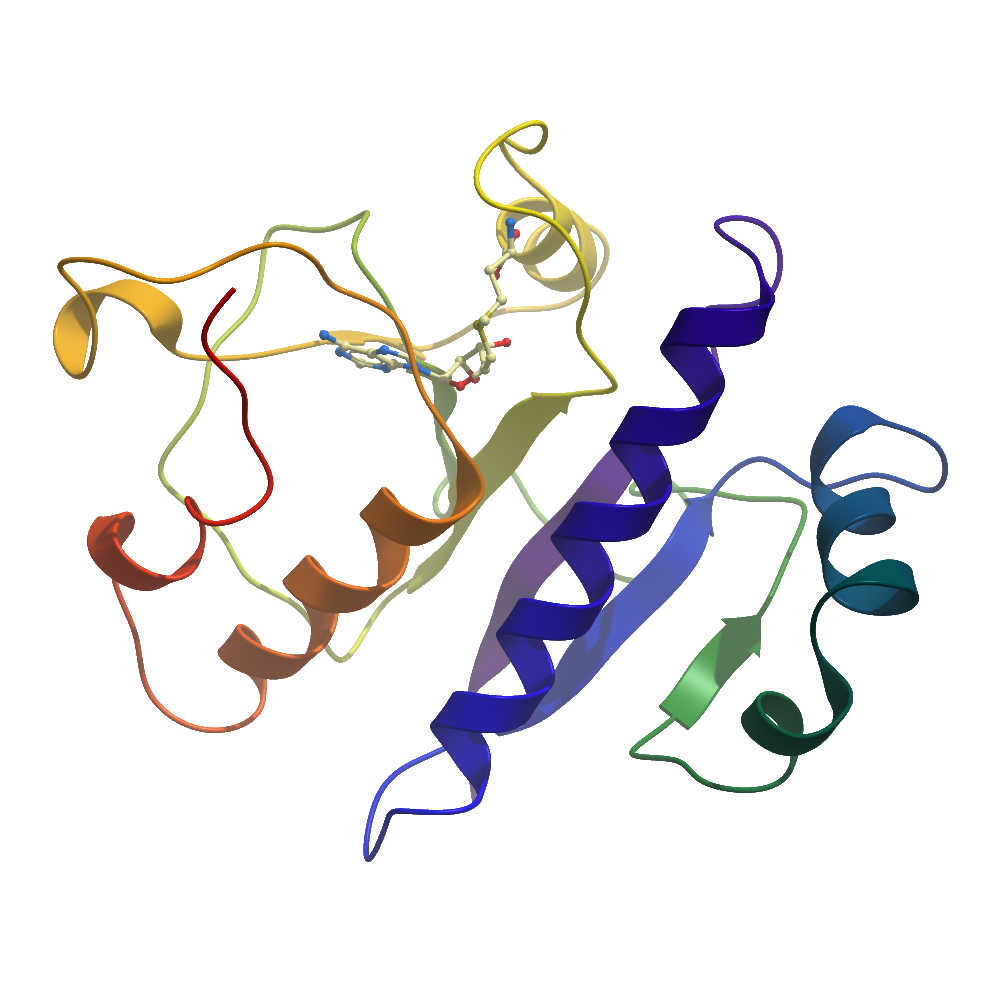Molecular Biology
Entry Clone Accession: IMAGE:4719195
Entry Clone Source: MGC
SGC Construct ID: RG9MTD1A-c113
Protein Region: Q203-T403
Vector: pNIC-CTHF
Tag: C-TEV;C-6HIS
Host: BL21(DE3)-R3-pRARE2
Sequence (with tag(s)): MQPLVFDMAYENYMKRKELQNTVSQLLESEGWNRRNVDPFHIYFCNLKIDGALHRELVKRYQEKWDKLLLTSTEKSHVDLFPKDSIIYLTADSPNVMTTFRHDKVYVIGSFVDKSMQPGTSLAKAKRLNLATECLPLDKYLQWEIGNKNLTLDQMIRILLCLKNNGNWQEALQFVPKRKHTGFLEISQHSQEFINRLKKAKTAENLYFQSHHHHHHDYKDDDDK
Sequence after tag cleavage: MQPLVFDMAYENYMKRKELQNTVSQLLESEGWNRRNVDPFHIYFCNLKIDGALHRELVKRYQEKWDKLLLTSTEKSHVDLFPKDSIIYLTADSPNVMTTFRHDKVYVIGSFVDKSMQPGTSLAKAKRLNLATECLPLDKYLQWEIGNKNLTLDQMIRILLCLKNNGNWQEALQFVPKRKHTGFLEISQHSQEFINRLKKAKTAENLYFQ
DNA Sequence: CTTAAGAAGGAGATATACTATGCAACCTTTGGTTTTTGACATGGCTTACGAAAATTATATGAAACGAAAAGAATTGCAGAATACTGTTTCCCAGCTTTTAGAAAGTGAAGGATGGAACAGAAGAAATGTTGATCCTTTCCATATTTATTTCTGCAATCTAAAAATAGATGGTGCTTTGCACAGAGAGTTAGTTAAACGGTATCAAGAAAAATGGGACAAATTGCTTTTAACATCAACAGAAAAGTCTCATGTAGATTTATTTCCAAAGGACAGTATTATCTATTTAACTGCAGATTCTCCCAATGTTATGACTACTTTCAGGCATGACAAAGTTTATGTAATTGGGTCTTTTGTTGATAAGAGTATGCAGCCAGGCACATCCCTAGCCAAGGCAAAACGGCTGAACCTGGCAACTGAATGCCTTCCATTAGATAAATATTTACAATGGGAAATTGGTAACAAAAATCTCACCTTAGATCAAATGATACGTATTTTGTTATGTCTGAAAAACAATGGTAATTGGCAAGAGGCTCTGCAATTCGTTCCCAAGAGAAAACATACTGGTTTTCTGGAGATTTCTCAGCATTCTCAAGAGTTTATCAACAGACTAAAGAAGGCAAAGACTGCAGAGAACCTCTACTTCCAATCGCACCATCATCACCACCATGATTACAAGGATGACGACGATAAGTGAGGATCC
Protein Expression
Medium: Terrific Broth
Antibiotics: Kanamycin
Procedure: Plasmids were transformed into E. coli BL21(DE3)-R3-pRARE2 competent cells, cultured in Terrific Broth at 37°C, and induced with 0.1 mM isopropyl β-D-1-thiogalactopyranoside for overnight growth at 18oC.
Protein Purification
Buffers:
Lysis buffer: 500 mM NaCl, 50 mM HEPES pH 7.5, 5% glycerol, 0.5 mM TCEP, 20 mM imidazole
Wash buffer: 500 mM NaCl, 50 mM HEPES pH 7.5, 5% glycerol, 0.5 mM TCEP, 40 mM imidazole
Elution buffer: 500 mM NaCl, 50 mM HEPES pH 7.5, 5% glycerol, 0.5 mM TCEP, 250 mM imidazole
SEC buffer: 500 mM NaCl, 50 mM HEPES pH 7.5, 5% glycerol, 0.5 mM TCEP
Procedure:
Cell lysis
Frozen cell pellets were thawed and resuspended on ice in 50 mL lysis buffer containing 1 mL protease inhibitor cocktail set III (Calbiochem) per L of harvested cell culture. The cells were lysed by passing five times on a homogeniser (EmulsiFlex-C5-Avestin) to a pressure of 1000 bar. The cell lysate was clarified by centrifugation (Beckman Coulter Avanti JXN-26) at 4ᵒC for 1 h at 16500 rpm.
Immobilised metal affinity chromatography (IMAC)
Clarified lysate from large-scale extraction was added with 2 mL of Ni-NTA affinity resin (GE Healthcare) and incubated at 4ᵒC for 1 h rotating at 8 rpm. The lysate, together with Ni-NTA resin, was centrifuged (Beckman Coulter Avanti JXN-26) for 30 min at 5000 rpm at 4ᵒC. The Ni-NTA pellet was applied onto a 1.5x10 cm (DxH) column (Bio-Rad) under gravity flow and the flow-through was collected for SDS-PAGE analysis. The Ni-NTA resin was washed with 2 x 10 CVs Lysis buffer and 2 x 10 CVs Wash buffer. The bound protein was eluted with 5 x 2 CVs of Elution buffer.
Size exclusion chromatography (SEC)
Pooled fractions from IMAC were concentrated to 5 mL using an Amicon Ultra centrifugal filter device (Merck Millipore) with a 10 kDa cut-off. The concentrated sample was loaded onto a 120 mL Sephadex 75 HiLoad 16/60 column connected to an AKTA-Express (GE Healthcare) in 4ᵒC. Before loading the protein, the column was pre-equilibrated with SEC buffer. Protein was loaded and eluted at a rate of 1 mL/min and 1.8 mL fractions were collected throughout the run.
Treatment with Tobacco etch virus protease
To remove the affinity tag from target proteins, Tobacco etch virus (TEV) protease was added to the eluates in a 1:20 (w/v) protease-to-protein ratio and the mixture was incubated overnight at 4ᵒC.
Reverse IMAC
TEV protease-treated sample was applied to a 2x10 cm gravity column containing 125 μL Ni-NTA affinity resin (GE Healthcare) equilibrated with Lysis buffer. The Ni-NTA resin was washed with 2 x 8 CVs Binding buffer and 2 x 8 CVs Wash buffer. Fractions containing protein were pooled.
Concentration: 10.93 mg/ml
Mass-spec Verification: Expected mass of the construct confirmed
Structure Determination
Crystallization: Crystals of hMRPP1MT were grown by vapour diffusion at 4°C, from sitting drops mixing 200 nL of protein (11 mg/ml; pre-incubated with 4.8 μM SAM) and 100 nL of reservoir solution containing either 14% PEG 1000, 28% glycerol or 12% PEG 1000, 28% glycerol, 1.5%(w/v) PEG 3350. Crystals were cryo-protected with reservoir solution supplemented with 25% (v/v) propylene glycol, and flash-cooled in liquid nitrogen. Heavy atom-derivatised crystals were obtained by soaking with 10mM C9H9HgNaO2S for 1h10min.
Data Collection: Diffraction data were collected on the Diamond Light Source beamline I04-1 for the HA-soaked crystals to 2.49 Å, and for two crystals of the as-purified protein to 2.22 Å and 1.96 Å.
Data Processing: Phases were determined by single-wavelength anomalous dispersion (SAD) with the SHELX package using the HA dataset. The phases from the HA-dataset were combined with the 2.22 Å native dataset, and the model was built in COOT. This model was used as search model in a molecular replacement done in PHASER from the CCP4 Suite using the native data collected to 1.96 Å. Modelling and refinement were carried out using Refmac and COOT.