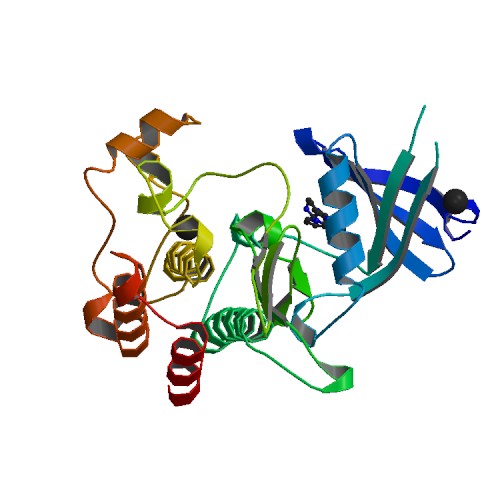
Molecular Biology
Entry Clone Accession: n/a
Entry Clone Source: Site-directed mutagenesis
SGC Construct ID: ACVR1A-c096
Protein Region: I208-D499
Vector: pFB-LIC-Bse
Tag: N-6HIS;N-TEV
Host: DH10Bac
Sequence (with tag(s)): MGHHHHHHSSGVDLGTENLYFQSMQRTVARDITLLECVGKGRYGEVWRGSWQGENVAVKIFSSRDEKSWFRETELYNTVMLRHENILGFIASDMTSRHSSTQLWLITHYHEMGSLYDYLQLTTLDTVSCLRIVLSIASGLAHLHIEIFGTQGKPAIAHRDLKSKNILVKKNGQCCIADLGLAVMHSQSTNQLDVGNNPRVGTKRYMAPEVLDETIQVDCFDSYKRVDIWAFGLVLWEVARRMVSNGIVEDYKPPFYDVVPNDPSFEDMRKVVCVDQQRPNIPNRWFSDPTLTSLAKLMKECWYQNPSARLTALRIKKTLTKID
Sequence after tag cleavage: SMQRTVARDITLLECVGKGRYGEVWRGSWQGENVAVKIFSSRDEKSWFRETELYNTVMLRHENILGFIASDMTSRHSSTQLWLITHYHEMGSLYDYLQLTTLDTVSCLRIVLSIASGLAHLHIEIFGTQGKPAIAHRDLKSKNILVKKNGQCCIADLGLAVMHSQSTNQLDVGNNPRVGTKRYMAPEVLDETIQVDCFDSYKRVDIWAFGLVLWEVARRMVSNGIVEDYKPPFYDVVPNDPSFEDMRKVVCVDQQRPNIPNRWFSDPTLTSLAKLMKECWYQNPSARLTALRIKKTLTKID
DNA Sequence: CCATGGGCCACCATCATCATCATCATTCTTCTGGTGTAGATCTGGGTACCGAGAACCTGTACTTCCAATCCATGCAAAGAACAGTGGCTCGCGATATTACACTGTTGGAGTGTGTCGGGAAAGGCAGGTATGGTGAGGTGTGGAGGGGCAGCTGGCAAGGGGAAAATGTTGCCGTGAAGATCTTCTCCTCCCGTGATGAGAAGTCATGGTTCAGGGAAACGGAATTGTACAACACTGTGATGCTGAGGCATGAAAATATCTTAGGTTTCATTGCTTCAGACATGACATCAAGACACTCCAGTACCCAGCTGTGGTTAATTACACATTATCATGAAATGGGATCGTTGTACGACTATCTTCAGCTTACTACTCTGGATACAGTTAGCTGCCTTCGAATAGTGCTGTCCATAGCTAGTGGTCTTGCACATTTGCACATAGAGATATTTGGGACCCAAGGGAAACCAGCCATTGCCCATCGAGATTTAAAGAGCAAAAATATTCTGGTTAAGAAGAATGGACAGTGTTGCATAGCAGATTTGGGCCTGGCAGTCATGCATTCCCAGAGCACCAATCAGCTTGATGTGGGGAACAATCCCCGTGTGGGCACCAAGCGCTACATGGCCCCCGAAGTTCTAGATGAAACCATCCAGGTGGATTGTTTCGATTCTTATAAAAGGGTCGATATTTGGGCCTTTGGACTTGTTTTGTGGGAAGTGGCCAGGCGGATGGTGAGCAATGGTATAGTGGAGGATTACAAGCCACCGTTCTACGATGTGGTTCCCAATGACCCAAGTTTTGAAGATATGAGGAAGGTAGTCTGTGTGGATCAACAAAGGCCAAACATACCCAACAGATGGTTCTCAGACCCGACATTAACCTCTCTGGCCAAGCTAATGAAAGAATGCTGGTATCAAAATCCATCCGCAAGACTCACAGCACTGCGTATCAAAAAGACTTTGACCAAAATTGATTGACAGTAAAGGTGGATACGGATCCGAATTCGAGCTCCGTCGACAAGCTT
Protein Expression
Medium: Insect Xpress
Antibiotics: Ampicillin
Procedure: Sf9 cells at a density of 2x106/ml were infected with recombinant ACVR1 baculovirus (virus stock P2; 3ml of virus stock/1000 ml of cell culture). Cells were shaken at 110 rpm at 27 °C in an Innova shaker. After 72 hours post-infection the cultures were harvested by centrifugation for 20min at 900rpm. Cell pellets from each 1L flask were resuspended in 15 ml binding buffer (50 mM Hepes, pH 7.5; 500 mM NaCl; 5% Glycerol; 5 mM imidazole). Calbiochem protease inhibitor SET V was added to the cell suspension at a 1:2000 dilution and transferred to 50 ml tubes, and stored at -20 °C.
Protein Purification
Procedure: The frozen cells were thawed and the volume increased to 160 ml with binding buffer. The cells were lysed by sonication over 12 min with the sonicator pulsing ON for 5 sec and OFF for 10 sec. The DNA was precipitated using 0.15% PEI (polyethyleneimine) pH 8. The cell lysate was spun down by centrifugation at 21.5K rpm at 4°C for 1 h. The supernatant was recovered for purification. Column 1: Ni-Affinity Chromatography. 6 ml of 50 % Ni-sepharose slurry was applied onto a 1.5 x 10 cm column. The column was equilibrated with binding buffer (25ml). Buffers: Binding buffer: 50 mM Hepes, pH 7.5; 500 mM NaCl; 5% Glycerol; 5 mM imidazole, 0.1mM TCEP Wash buffer: 50 mM Hepes, pH 7.5; 500 mM NaCl; 5% Glycerol; 25 mM imidazole, 0.1mM TCEPElution buffer: 50 mM HEPES, pH 7.5; 500 mM NaCl; 5% Glycerol; 50 to 250 mM imidazole, 0.1mM TCEP Procedure: The supernatant was applied by gravity flow onto the Ni-sepharose column. The bound protein was eluted by applying a step gradient of imidazole using 10 ml portions of elution buffer with increasing concentration of imidazole (50 mM, 100 mM, 150 mM, 250 mM). Enzymatic treatment: 0.1mg of TEV protease was added to the Ni-eluted protein (Overnight incubation at 4 °C) to remove the tag. Column 2: Size Exclusion Chromatography S200 HiLoad 16/60 Superdex run on ÄKTA-Express Buffer: Gel Filtration buffer: 300 mM NaCl, 50 mM Hepes pH 7.5, 0.5mM TCEP Procedure: Prior to applying the protein, the S200 16/60 column was washed and equilibrated with gel filtration buffer. The protein was concentrated to 4 ml using an Amicon Ultra-15 filter with a 10 kDa cut-off. The concentrated protein was directly applied onto the equilibrated S200 16/60 column, and run at a flow-rate of 1 ml/min. The protein was eluted at 90-110 ml. Fractions containing the protein were pooled together and stored with 10mM DTT. Column 3: Ni-rebinding. 2 ml of 50% Ni sepharose slurry was applied to a 1.5 x 10 cm column. The column was equilibrated with binding buffer (25ml). Buffers: Binding buffer: 50 mM Hepes, pH 7.5; 500 mM NaCl; 5% Glycerol; 5 mM imidazole, 0.1mM TCEP Wash buffer: 50 mM Hepes, pH 7.5; 500 mM NaCl; 5% Glycerol; 25 mM imidazole, 0.1mM TCEPElution buffer: 50 mM HEPES, pH 7.5; 500 mM NaCl; 5% Glycerol; 50 to 250 mM imidazole, 0.1mM TCEP. Procedure: The sample was diluted to reduce DTT to 2mM and applied by gravity flow onto the Ni-sepharose column. The flow through was collected containing the cleaved protein. The column was washed and any uncleaved or contaminating protein attached to the column was eluted in 250 mM imidazole buffer. The protein in the flow through was found to be pure and concentrated to 13.6 mg/ml in the presence of 50 mM Arg/Glu for crystallization.
Columns: Column 1: Ni Drip Column; Column 2: s200 16/60; Column 3: Ni-rebinding;
Concentration: 13.6 mg/ml
Mass-spec Verification: The purified protein was homogeneous and had an experimental mass of 34.492 kDa, as expected from primary sequences. Mass was determined by LC-MS, using an Agilent LC/MSD TOF system with reversed-phase HPLC coupled to electrospray ionisation and an orthogonal time-of-flight mass analyser. Proteins were desalted prior to mass spectrometry by rapid elution off a C3 column with a gradient of 5-95% acetonitrile in water with 0.1% formic acid.
Compound Exact Mass: 406.190595
Structure Determination
Crystallization: Protein was buffered in 50 mM HEPES pH 7.5, 300 mM NaCl, 2 mM DTT and 50mM L-arginine, 50 mM L-glutamate. The protein was concentrated to 13.6 mg/ml (calculated using an extinction co-efficient of 58900) in the presence of the inhibitor K03848b (LDN-212854) (1 mM end concentration). Crystals were grown at 4 °C in 150 nl sitting drops mixing 100 nl protein solution with 50 nl of a reservoir solution containing 18% PEG8000, 0.2M calcium acetate, 0.1M cacodylate pH6.5. On mounting crystals were cryoprotected with mother liquor plus 25% ethylene glycol before transfer to liquid nitrogen.
Data Collection: Diffraction data were collected at the Diamond Light Source, station I03 using monochromatic radiation at wavelength 0.97625 Å. Data could be analysed to a resolution of 1.73 Å.
Data Processing: Data were processed with MOSFLM [1] and subsequently scaled using the program AIMLESS from the CCP4 suite. Initial phases were obtained by molecular replacement using the program PHASER and the structure of ALK2 (Protein Data Bank code 3H9R) as a search model. The resulting structure solution was refined using REFMAC5 from the CCP4 suite, and manually rebuilt with COOT. Appropriate TLS restrained refinement using the tls tensor files calculated from the program TLSMD was applied at the final round of refinement. The complete structure was verified for geometric correctness with MolProbity.