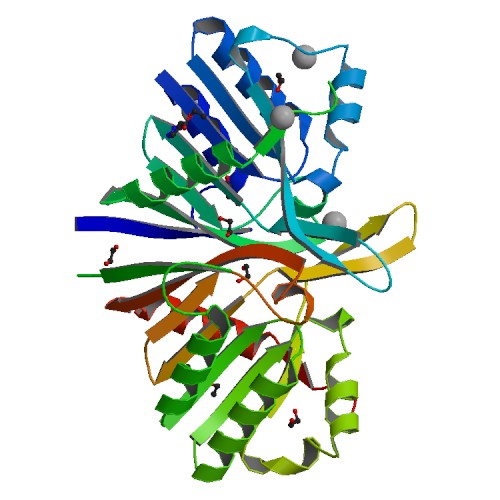
Molecular Biology
Entry Clone Accession: BC101628
Entry Clone Source: MGC
SGC Construct ID: FAM83BA-c004
Protein Region: G117-V294
Vector: pNIC28-Bsa4
Tag: N-6HIS;N-TEV
Host: BL21(DE3)-R3-pRARE2
Sequence (with tag(s)): MHHHHHHSSGVDLGTENLYFQSMGGTHIDLLFHPPRAHLLTIKETIRKMIKEARKVIALVMDIFTDVDIFKEIVEASTRGVSVYILLDESNFNHFLNMTEKQGCSVQRLRNIRVRTVKGQDYLSKTGAKFHGKMEQKFLLVDCQKVMYGSYSYMWSFEKAHLSMVQIITGQLVESFDEEFRTLYARSCVPSSFAQEESARV
Sequence after tag cleavage: SMGGTHIDLLFHPPRAHLLTIKETIRKMIKEARKVIALVMDIFTDVDIFKEIVEASTRGVSVYILLDESNFNHFLNMTEKQGCSVQRLRNIRVRTVKGQDYLSKTGAKFHGKMEQKFLLVDCQKVMYGSYSYMWSFEKAHLSMVQIITGQLVESFDEEFRTLYARSCVPSSFAQEESARV
DNA Sequence: CATATGCACCATCATCATCATCATTCTTCTGGTGTAGATCTGGGTACCGAGAACCTGTACTTCCAATCCATGGGGGGCACCCATATAGATCTCCTTTTTCATCCACCAAGAGCACATCTACTTACGATAAAAGAAACTATTCGGAAGATGATAAAAGAAGCAAGAAAGGTCATTGCTTTAGTGATGGATATATTTACAGATGTGGACATTTTCAAAGAAATCGTTGAGGCATCAACTCGAGGAGTATCTGTTTACATTCTGCTTGATGAGTCCAATTTTAATCATTTTCTAAATATGACTGAGAAACAAGGTTGTTCAGTTCAGCGTCTCAGGAATATTCGAGTGCGAACAGTAAAAGGCCAAGATTATCTTTCAAAAACAGGGGCAAAATTCCATGGAAAAATGGAACAGAAATTTTTGTTAGTTGACTGCCAGAAAGTGATGTACGGTTCTTACAGTTATATGTGGTCATTTGAGAAAGCTCACCTCAGCATGGTTCAGATAATTACAGGACAACTTGTTGAGTCCTTTGATGAAGAATTTAGAACTCTCTATGCCAGATCCTGTGTCCCTAGTTCATTTGCTCAGGAAGAATCAGCAAGGGTGTGACAGTAAAGGTGGATACGGATCCGAA
Protein Expression
Medium: LB
Antibiotics: Kanamycin/Chloramphenicol
Procedure: A glycerol stock was used to inoculate a starter culture which was supplemented with Kanamycin (50 µg/ml)/ Chloramphenicol (34 µg/ml). These were then grown overnight at 37 °C with shaking at 240 RPM. The starter culture was used to inoculate 4 x 1 litre LB flasks supplemented with Kanamycin/Chloramphenicol. The cultures were then grown at 37 °C with shaking at 160 RPM to an OD600 of between 0.6-0.8. The cultures were then cooled to 18 °C, induced with 0.35 mM IPTG and the protein was expressed overnight at 18 °C /160 RPM. Cultures were harvested, pellets made up to 40 ml with Nickel affinity binding buffer (50 mM HEPES pH 7.5, 500 mM NaCl, 5 mM Imidazole, 5% Glycerol) and 40 ul of SET III protease inhibitors (Merck) added before storage at -20 °C.
Protein Purification
Procedure: Cells were lysed by sonication, 0.125% polyethyleneimine added, and lysates were clarified by centrifugation at 50,000G. The supernatant was then applied to a Nickel-NTA gravity column and washed and eluted with 50 mM HEPES pH 7.5, 500 mM NaCl, 5% glycerol, and 10-300mM imidazole. Protein containing fractions were then treated with TEV protease overnight at 4 °C prior to gel filtration on an S200 gel filtration column using 50 mM HEPES pH 7.5, 300 mM NaCl, 0.5 mM TCEP as the running buffer. The protein was additionally purified by cation exchange using a HiTrap S column and eluted using increasing salt concentrations from 50 mM NaCl to 1000 mM NaCl in 50 mM HEPES pH 7.5, 0.5 mM TCEP. Protein eluted at approximately 200 mM NaCl and was concentrated to 18.4 mg/ml using a spin concentrator.
Columns: Column 1: Ni-NTA; Column 2: S200 gel filtration; Column 3: HiTrap S cation exchange
Concentration: 18.4 mg/ml
Mass-spec Verification: Intact mass correct
Structure Determination
Crystallization: Crystals were grown using the sitting drop vapor diffusion method at 4 ֯C. Crystals were grown in 300 nl drops consisting of 2:1 mother liquor (30.5% PEG3350 -- 0.2M sodium iodide -- 0.1M bis-tris-propane pH 9.1 -- 10% ethylene glycol) to protein (12.0 mg/ml). To the crystallization drop, 10% of the crystallization drop volume (30 nl) of 500 mM fragment (in this case N13653a a.k.a. ZINC00409316 a.k.a. FMOPL000271a) in DMSO was added, a soaked crystal was then mounted and flash frozen in liquid nitrogen without additional cryoprotection.
Data Collection: Data was collected at beamline I04-1 at 100K.
Data Processing: Data was processed to a resolution of approximately 1.7 Å using XDS and either xia2 or autoPROC, or DIALS.