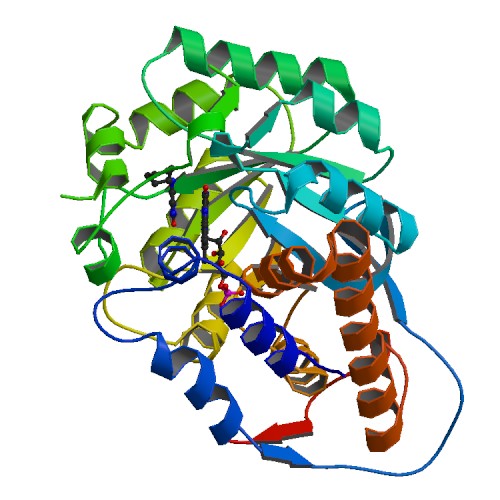
Molecular Biology
Entry Clone Accession: NM_017545
Entry Clone Source: Origene
SGC Construct ID: HAO1A-c002
Protein Region: M1-S368
Vector: pNIC28-Bsa4
Tag: N-6HIS; TEV cleavage site
Host: BL21(DE3)-R3-pRARE2
Sequence (with tag(s)): MHHHHHHSSGVDLGTENLYFQSMLPRLICINDYEQHAKSVLPKSIYDYYRSGANDEETLADNIAAFSRWKLYPRMLRNVAETDLSTSVLGQRVSMPICVGATAMQRMAHVDGELATVRACQSLGTGMMLSSWATSSIEEVAEAGPEALRWLQLYIYKDREVTKKLVRQAEKMGYKAIFVTVDTPYLGNRLDDVRNRFKLPPQLRMKNFETSTLSFSPEENFGDDSGLAAYVAKAIDPSISWEDIKWLRRLTSLPIVAKGILRGDDAREAVKHGLNGILVSNHGARQLDGVPATIDVLPEIVEAVEGKVEVFLDGGVRKGTDVLKALALGAKAVFVGRPIVWGLAFQGEKGVQDVLEILKEEFRLAMALSGCQNVKVIDKTLVRKNPLAVS
Sequence after tag cleavage: SMLPRLICINDYEQHAKSVLPKSIYDYYRSGANDEETLADNIAAFSRWKLYPRMLRNVAETDLSTSVLGQRVSMPICVGATAMQRMAHVDGELATVRACQSLGTGMMLSSWATSSIEEVAEAGPEALRWLQLYIYKDREVTKKLVRQAEKMGYKAIFVTVDTPYLGNRLDDVRNRFKLPPQLRMKNFETSTLSFSPEENFGDDSGLAAYVAKAIDPSISWEDIKWLRRLTSLPIVAKGILRGDDAREAVKHGLNGILVSNHGARQLDGVPATIDVLPEIVEAVEGKVEVFLDGGVRKGTDVLKALALGAKAVFVGRPIVWGLAFQGEKGVQDVLEILKEEFRLAMALSGCQNVKVIDKTLVRKNPLAVS
DNA sequence: CATATGCACCATCATCATCATCATTCTTCTGGTGTAGATCTGGGTACCGAGAACCTGTACTTCCAATCCATGCTCCCCCGGCTAATTTGTATCAATGATTATGAACAACATGCTAAATCAGTACTTCCAAAGTCTATATATGACTATTACAGGTCTGGGGCAAATGATGAAGAAACTTTGGCTGATAATATTGCAGCATTTTCCAGATGGAAGCTGTATCCAAGGATGCTCCGGAATGTTGCTGAAACAGATCTGTCGACTTCTGTTTTAGGACAGAGGGTCAGCATGCCAATATGTGTGGGGGCTACGGCCATGCAGCGCATGGCTCATGTGGACGGCGAGCTTGCCACTGTGAGAGCCTGTCAGTCCCTGGGAACGGGCATGATGTTGAGTTCCTGGGCCACCTCCTCAATTGAAGAAGTGGCGGAAGCTGGTCCTGAGGCACTTCGTTGGCTGCAACTGTATATCTACAAGGACCGAGAAGTCACCAAGAAGCTAGTGCGGCAGGCAGAGAAGATGGGCTACAAGGCCATATTTGTGACAGTGGACACACCTTACCTGGGCAACCGTCTGGATGATGTGCGTAACAGATTCAAACTGCCGCCACAACTCAGGATGAAAAATTTTGAAACCAGTACTTTATCATTTTCTCCTGAGGAAAATTTTGGAGACGACAGTGGACTTGCTGCATATGTGGCTAAAGCAATAGACCCATCTATCAGCTGGGAAGATATCAAATGGCTGAGAAGACTGACATCATTGCCAATTGTTGCAAAGGGCATTTTGAGAGGTGATGATGCCAGGGAGGCTGTTAAACATGGCTTGAATGGGATCTTGGTGTCGAATCATGGGGCTCGACAACTCGATGGGGTGCCAGCCACTATTGATGTTCTGCCAGAAATTGTGGAGGCTGTGGAAGGGAAGGTGGAAGTCTTCCTGGACGGGGGTGTGCGGAAAGGCACTGATGTTCTGAAAGCTCTGGCTCTTGGCGCCAAGGCTGTGTTTGTGGGGAGACCAATCGTTTGGGGCTTAGCTTTCCAGGGGGAGAAAGGTGTTCAAGATGTCCTCGAGATACTAAAGGAAGAATTCCGGTTGGCCATGGCTCTGAGTGGGTGCCAGAATGTGAAAGTCATCGACAAGACATTGGTGAGGAAAAATCCTTTGGCCGTTTCCTGACAGTAAAGGTGGATACGGATCCGAA
Protein Expression
Medium: Terrific Broth
Antibiotics: Kanamycin
Procedure: An overnight culture (20 mL LB) was used to inoculate 2L auto-induction TB containing 50 µg/ml each of kanamycin and chloramphenicol. Cells were cultured at 37°C for 6 hours followed by 48 h incubation at 18°C.
Protein Purification
Procedure: Cell pellet from 2 L of culture was re-suspended in 150 mL Extraction Buffer, lysed by sonication for 15 minutes, and centrifuged at 37000 x g for 1 hour at 4°C. The clarified cell extract was incubated with 5 mL of Ni-NTA resin pre-equilibrated with lysis buffer before applying to 1.5 x 10 cm column by gravity flow. The column was washed with 10 column volumes of Binding Buffer, 10 column volumes of Wash Buffer, and eluted with 5 x 5 mL Elution Buffer. Fractions containing hHAO1 were loaded onto gel filtration column (Superdex 200 Hiload 16/60) pre-equilibrated with GF buffer. Fractions containing hHAO1 were pooled.
Extraction Buffer: 500 mM NaCl, 50 mM HEPES pH 7.5, 20 mM imidazole, 0.5 mM TCEP, 5% glycerol, 1:1000 of Merck Protease Cocktail II, 0.5 mg/mL lysozyme and 0.2 µg/mL benzonase.
Binding Buffer: 500 mM NaCl, 50 mM HEPES pH 7.5, 20 mM imidazole, 0.5 mM TCEP, 5% glycerol.
Washing Buffer: 500 mM NaCl, 50 mM HEPES pH 7.5, 40 mM imidazole, 0.5 mM TCEP, 5% glycerol.
Elution Buffer: 500 mM NaCl, 50 mM HEPES pH 7.5, 250 mM imidazole, 0.5 mM TCEP, 5% glycerol.
GF buffer: 500 mM NaCl, 550 mM HEPES pH 7.5, 0.5 mM TCEP, 5% glycerol.
Concentration: 13.7 mg/ml
Mass-spec Verification: Yes
Structure Determination
Crystallization: Crystals were grown by vapour diffusion at 4°C in sitting drops of 13.7 mg/mL protein equilibrated against well solution containing 25-35% PEG1000, sodium malonate-imidazole-boric acid pH 8.0. Approximately 50 crystals of suitable size were identified per 96-well crystallisation plate, and a total of 10 plates generated sufficient crystals for the entire fragment campaign.
Data Collection: Crystals were soaked for two hours with fragments from the Diamond-SGC Poised Library before being harvested using XChem SHIFTER technology, cryo-cooled in liquid nitrogen, and data sets collected at the beamline I04-1 in “automated unattended” mode.
Data Processing: The XChemXplorer pipeline was used for structure solution with parallel molecular replacement using DIMPLE (template 2NZL), followed by map averaging and statistical modelling to identify weak electron densities generated from low occupancy fragments using Pandda software.