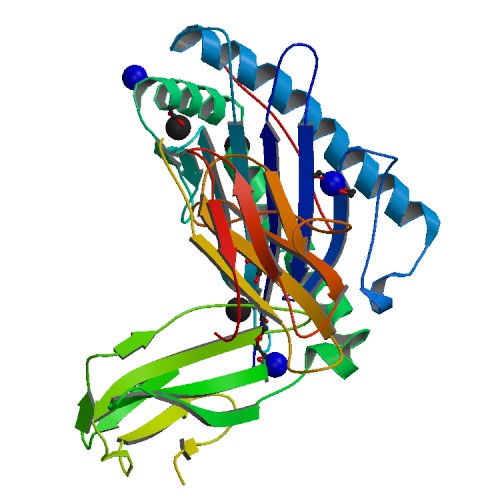
Molecular Biology
Entry Clone Accession:
Entry Clone Source: David Gfeller
SGC Construct ID: B2MA-c001
Protein Region: I21-M119
Vector: pHN1+
Tag: no tag
Host: XA-90
Sequence (with tag(s)): MIQRTPKIQVYSRHPAENGKSNFLNCYVSGFHPSDIEVDLLKNGERIEKVEHSDLSFSKDWSFYLLYYTEFTPTEKDEYACRVNHVTLSQPKIVKWDRDM
DNA Sequence: CTTAAGAAGGAGATATACTATGATCCAGCGTACTCCAAAGATTCAGGTTTACTCACGTCATCCAGCAGAGAATGGAAAGTCAAATTTCCTGAATTGCTATGTGTCTGGGTTTCATCCATCCGACATTGAAGTTGACTTACTGAAGAATGGAGAGAGAATTGAAAAAGTGGAGCATTCAGACTTGTCTTTCAGCAAGGACTGGTCTTTCTATCTCTTGTACTACACTGAATTCACCCCCACTGAAAAAGATGAGTATGCCTGCCGTGTGAACCATGTGACTTTGTCACAGCCCAAGATAGTTAAGTGGGATCGAGACATGTGAGCAGAGAACCTCTACTTCCAATCGCACCATCATCACCACCATGATTACAAGGATGACGACGATAAGTGAGGATCC
Protein Expression
Medium: Luria-Bertani medium
Antibiotics: Kanamycin
Procedure: Recombinant expression plasmids were transformed into BL21 (DE3) (HLA-A68:01) or XA90 strain (β2M) bacteria. The cells were grown overnight in LB (Luria-Bertani medium) twice concentrated (2X) supplemented with 50 µg ml-1 kanamycin or 100 µg ml-1 ampicillin respectively at 37 °C. One litre of pre-warmed LB was inoculated with 10 ml of the overnight culture and was incubated at 37 °C. At an OD600 nm of 0.6, expression was induced for 8 h at 37 °C with 1.0 mM isopropyl-β-D-thiogalactopyranoside (IPTG). Cells were then harvested by centrifugation (8700xg, 15 min, 4 °C, Beckman Coulter Avanti J-20 XP centrifuge) and pellets were stored at -80°C
Protein Purification
Procedure: The heavy chain HLA-A68:01 and the light chain b2M were purified from inclusion-bodies. Cell pellets were re-suspended in lysis buffer (10 mM TRIS-HCl, pH 8.0 at 20 °C complemented with 100 μg ml-1 Lysozyme (Sigma Aldrich), 250 units of Benzonase (Novagen), 1 mM EDTA and 1:1000 (v/v) Protease Inhibitor Cocktail III (Calbiochem)). After 20 min of continuous rocking at 22 °C, cells were lysed 3 times at 4 °C using a Basic Z-Model Cell Disrupter (Constant Systems Ltd, UK). Inclusion-bodies were isolated by centrifugation (16000xg for 1 h at 4 °C, JA 25.50 rotor, on a Beckman Coulter Avanti J-20 XP centrifuge). The supernatant was decanted and the contaminated material was removed by scrapping the outer rings (viscous and dark coloured) leaving the more compact and lighter coloured inner ring (containing the protein of interest) intact. The pellet was then fully dispersed in 20 ml of washing buffer (10 mM TRIS-HCl, pH 8.0 at 20 °C) complemented with 1:1000 (v/v) Protease Inhibitor Cocktail III (Calbiochem) and centrifuged again for 10 min (16000xg at 4 °C, JA 25.50 rotor, on a Beckman Coulter Avanti J-20 XP centrifuge). The washing/scrapping steps were repeated 5 times (until the outer rings vanished). The pellet containing the recombinant protein was then dissolved in 20 ml of solubilisation buffer (100 mM TRIS-HCl, pH 8.0 at 20 °C, 8 M urea). Insoluble material was precipitated by centrifugation (16000xg for 1 h at 4 °C, JA 25.50 rotor, on a Beckman Coulter Avanti J-20 XP centrifuge). Solubilized HLA-A68:01 heavy chain was immediately flash frozen in liquid nitrogen and stored at -80 °C. The recombinant 2M protein in urea was refolded by dialysis against 10 mM TRIS-HCl pH7.0 using SnakeSkin® Dialysis tubing (3.500 MWCO, Thermo Scientific) and purified by ion exchange on Hi-Trap Q HP (5 ml, GE Healthcare) column, with a linear gradient from 0 to 100 mM NaCl. Fractions containing pure 2M were dialysed overnight against water, concentrated with a 10 MWCO concentrator (Amicon® Ultra, MILLIPORE) to 2 mg ml-1, flash frozen in liquid nitrogen and stored at -80 °C.
Complex Formation
The HLA:β2M:peptide complex was reconstituted by dilution of the denatured HLA-A68:01 heavy chain (3 μM) and β2M (6 μM) in presence of the peptide ([H]-E-T-S-P-L-T-A-E-K-L-[OH], 10 μM) into 200 ml of refolding buffer (100 mM TRIS-HCl , pH 8.0 at 20 °C, 400 mM L-Arginine HCl (Sigma Aldrich), 2 mM EDTA, 5 mM reduced L-glutathione (Sigma Aldrich), 0.5 mM oxidized L-glutathione (Sigma Aldrich) and with 1:1000 (v/v) Protease Inhibitor Cocktail III (Calbiochem)). The refolding mixture was incubated at 10 °C during 36 h under constant stirring. Every 12 h another batch of the denatured HLA-A68:01 heavy chain (3 μM) was added to the mix. The 200 ml were then concentrated to 5 ml using a 3 MWCO concentrator (Amicon® Ultra, MILLIPORE). The concentrated protein mixture was further submitted to exclusion size chromatography (HiLoad™ 16/60 Superdex™ 75 prep grade, on Äkta Pure GE Healthcare) and each peak collected was submitted to SDS-PAGE gel and mass spectrometry (Agilent 6530 QTOF (Agilent Technologies Inc. - Palo Alto, CA)) to assess the simultaneous presence of the 3 components of the complex. Fractions of interest were then stored at 4°C until further use.
Concentration: 1.6 mg/ml
Mass-spec Verification: 11863.24
Compound Exact Mass:11862.36
Structure Determination
Crystallization: Prior to crystallization, the buffer of the protein complex was exchanged to 25 mM MES pH6.5 at 20 °C and 150 mM NaCl on a Superdex™ 200 Increase 10/300 GL column using an Äkta Pure system (GE Healthcare). The complex was then concentrated to 10.96 mg ml-1 final using a 3 kDa MWCO concentrator (Amicon® Ultra, MILLIPORE). Aliquots of the complex were set up for crystallization using a mosquito® crystallization robot (TTP Labtech, Royston UK). Coarse screens were typically setup onto Greiner 3-well plates using three different drop ratios of precipitant to protein per condition (100+50 nl, 75+75 nl and 50+100 nl). Crystallization was carried out using the sitting drop vapor diffusion method at 4 °C. Crystals were grown by mixing 100 nl of the protein complex (10.96 mg/ml) with 50 nl of reservoir solution containing 0.1 M HEPES pH 7.5, 12 % PEG3350, 0.005 M CoCl2, 0.005 M NiCl2, 0.005 M CdCl2 and 0.005 M MgCl2. Diffraction quality crystals grew within a few days.
Data Collection: Crystals were cryo-protected using the well solution supplemented with additional ethylene glycol and were flash frozen in liquid nitrogen. Data were collected at Diamond beamline I24 on a Pilatus3 6M detector at a wavelength of 0.96864
Data Processing: Indexing and integration was carried out using XDS (Kabsch, 2010) and scaling was performed with SCALA (Evans, 2017). Initial phases were calculated by molecular replacement with PHASER (McCoy et al., 2005) using a model of HLA/B2MG peptide complex (PDB ID 5T6X). Initial models were built by ARP/wARP (Perrakis et al., 1999) followed by manual building in COOT (Emsley and Cowtan, 2004). Refinement was carried out in REFMAC5 (Murshudov et al., 1997). Thermal motions were analyzed using TLSMD (Painter and Merritt, 2006) and hydrogen atoms were included in late refinement cycles.